
Expert Insights and Updates in Scientific Consulting
Click on the links below, to access the specific articles:
|
|
|

My interview to Nature journal regarding be paid to provide peer reviews
Back to top
Space Study Reveals Accelerated Growth of Human Brain Cells
Back to top
Unlocking the Language of RNA: AI Model Revolutionizes Plant Science and Agriculture
In the realm of molecular biology, RNA plays a crucial role as the messenger and regulator of genetic information. Yet, decoding the intricate "language" of RNA has long been a challenge for scientists. Enter PlantRNA-FM, a groundbreaking AI model introduced in a recent publication in Nature Machine Intelligence. This innovative model has the potential to reshape our understanding of RNA and revolutionize agricultural science.
What Is PlantRNA-FM?
PlantRNA-FM is a foundation model trained on RNA data from over 1,100 plant species. Its purpose? To uncover patterns and motifs in RNA sequences that could lead to better comprehension of genetic regulation, plant communication, and even responses to environmental stress.
This AI model leverages cutting-edge computational techniques, including motif-aware pretraining, which captures both subsequence patterns and motif-level features in RNA. Additionally, a type‐guided fine‐tuning approach allows the model to adapt across diverse RNA‐related tasks, integrating predictions about RNA types to enhance its accuracy and utility.
Why Does This Matter?
RNA is at the core of many biological processes, yet our understanding of its functions ‐ especially in plants ‐has been limited. PlantRNA-FM breaks new ground by enabling researchers to:
1. Identify Functional RNA Elements: The model provides insights into how specific RNA sequences regulate gene expression, paving the way for breakthroughs in plant biology.
2. Understand Plant Communication: By decoding RNA‐based signaling, PlantRNA-FM could help uncover how plants respond to stressors like drought, pests, and climate change.
3. Advance Agricultural Innovation: With its ability to analyze RNA at an unprecedented scale, this tool could accelerate the development of more resilient and productive crops.
Key Findings
The study behind PlantRNA-FM demonstrated the model's versatility across a range of tasks, including RNA sequence classification, motif prediction, and functional annotation. It significantly outperformed existing tools in these areas, proving to be an invaluable resource for RNA research.
This versatility makes PlantRNA-FM not just a scientific breakthrough but also a tool with practical applications in agriculture and biotechnology. For instance, farmers and crop scientists could use insights from PlantRNA-FM to develop crops that are better equipped to withstand environmental changes, contributing to global food security.
Implications for the Future
As AI continues to make strides in biology, models like PlantRNA-FM highlight the power of machine learning in solving complex biological puzzles. By automating and enhancing the analysis of RNA sequences, PlantRNA-FM sets the stage for a new era in plant science ‐one where data-driven insights lead to tangible benefits for both ecosystems and agriculture.
This innovation doesn't just push the boundaries of what we know about RNA; it underscores the role of interdisciplinary approaches ‐ combining biology, computational science, and AI‐in addressing some of the world's most pressing challenges.
At SCI Consultoria Científica, we are always inspired by such transformative advancements. As leaders in fostering science communication, we believe in showcasing the power of research to drive innovation and societal progress. If you're interested in exploring more about cutting-edge discoveries like PlantRNA-FM, visit our blog.
Let's celebrate the power of science and technology to create a better future for all!

An interpretable RNA foundation model for exploring functional RNA motifs in plants
Back to top
Revolutionizing Molecular Biology: CRISPR 3.0 - The Next Frontier in Gene Editing
The field of molecular biology is no stranger to groundbreaking discoveries, and now we stand on the cusp of yet another revolutionary advancement: CRISPR 3.0, a next-generation gene-editing technology that promises unparalleled precision, efficiency, and potential applications.
For years, the original CRISPR-Cas9 system transformed genetic engineering by enabling scientists to edit DNA with unprecedented ease. Yet, like all technologies, CRISPR has evolved. CRISPR 3.0 builds on this legacy with several innovative enhancements that could redefine molecular biology and medicine as we know them.
What Makes CRISPR 3.0 Special?
1. Prime Editing for Greater Precision
Unlike traditional CRISPR, which relies on creating double-stranded breaks in DNA, CRISPR 3.0 incorporates prime editing. This allows scientists to ’┐Įsearch and replace’┐Į single nucleotides or larger segments of DNA without damaging the genetic material. It significantly reduces the risks of unintended mutations and makes it safer for therapeutic applications.
2. RNA-Targeting CRISPR
While previous versions primarily targeted DNA, CRISPR 3.0 introduces robust tools for editing RNA. This innovation opens the door for transient, reversible gene editing, ideal for diseases where permanent changes might carry risks, such as neurological disorders or cancer.
3. Base Editing for Silent Mutations
CRISPR 3.0 includes advanced base editors that can convert one nucleotide into another, targeting even the smallest ’┐Įsilent mutations’┐Į responsible for many genetic diseases. This capability holds the promise of curing disorders that were once deemed intractable.
4. Multifunctionality and Multiplexing
CRISPR 3.0 systems are designed to edit multiple genes simultaneously with high precision. This innovation is particularly exciting for tackling polygenic diseases, such as diabetes, cardiovascular disease, and many forms of cancer.
Real-World Applications
’┐Į Personalized Medicine: CRISPR 3.0 could enable tailored treatments for genetic disorders, allowing for individualized therapies that correct disease-causing mutations without off-target effects.
’┐Į Synthetic Biology: Scientists could engineer microbes to produce biofuels, biodegradable plastics, or even innovative drugs using enhanced gene-editing tools.<
’┐Į Agricultural Revolution: With the potential to edit plant and animal genomes with greater precision, CRISPR 3.0 could usher in a new era of sustainable agriculture, creating crops that are more resistant to climate change, pests, and diseases.
’┐Į Epidemic Response: The RNA-targeting capabilities of CRISPR 3.0 could serve as a rapid response tool for viral outbreaks, enabling real-time suppression of pathogenic RNA.
Challenges and Ethical Considerations
While the potential of CRISPR 3.0 is staggering, it comes with challenges. Scientists are working tirelessly to address off-target effects, delivery mechanisms, and scalability. Furthermore, ethical questions surrounding gene editing remain prominent, particularly regarding germline editing and its implications for future generations.
The Future of Science and Humanity
CRISPR 3.0 is more than a technical advancement; it is a testament to human ingenuity and the relentless pursuit of knowledge. As we refine this technology and expand its applications, the possibilities for improving life on Earth’┐Įand perhaps beyond’┐Įare limitless.
Join the Conversation: What do you think are the most exciting applications of CRISPR 3.0? How should the scientific community address its ethical implications? Share your thoughts and tag us on social media!
Visit our blog regularly for more updates on cutting-edge advancements in molecular biology and science. Together, let’┐Įs explore the frontiers of discovery!
AlphaFold 3: revolutionizing our understanding of biomolecular interactions
The latest breakthrough in computational biology, AlphaFold 3, represents a monumental step forward in the field of biomolecular structure prediction. Developed by DeepMind, AlphaFold 3 builds upon the remarkable achievements of its predecessor, AlphaFold 2, which gained worldwide recognition for accurately predicting protein folding. This new iteration takes structural biology to new heights by modeling the complex interactions between proteins, DNA, RNA, and other biomolecules.
AlphaFold 3 introduces a groundbreaking diffusion-based architecture, enabling unparalleled precision in predicting the joint structures of biomolecular assemblies. This capability is pivotal for unraveling the intricate mechanisms that govern biological processes. From cellular signaling pathways to the molecular basis of disease, the system offers critical insights that were previously out of reach. By modeling these interactions, AlphaFold 3 holds immense promise for identifying therapeutic targets and advancing drug discovery efforts.
What makes AlphaFold 3 even more impactful is its accessibility. As an open-access tool, it empowers researchers across the globe to tackle complex questions in biology, accelerating innovation in ways that were unimaginable just a few years ago. Whether it is understanding the mechanisms of genetic disorders or exploring new avenues in biotechnology, AlphaFold 3 levels the playing field for scientists everywhere.
This powerful convergence of artificial intelligence and molecular biology exemplifies the transformative potential of interdisciplinary collaboration. AlphaFold 3 not only pushes the boundaries of what we know about biomolecular interactions but also sets the stage for rapid advancements in biomedical research. As we continue to harness AI to decode the complexities of life, tools like AlphaFold 3 are shaping a future where scientific discovery and innovation thrive at an unprecedented pace.

More could be read at:Accurate structure prediction of biomolecular interactions with AlphaFold 3
Stay tuned for more updates on how cutting-edge technologies are revolutionizing science at SCICC Blog.
#artificialintelligence #molecularbiology #AlphaFold3 #scientificbreakthrough #drugdiscovery
Back to top
In a landmark discovery, researchers from the University of California, Irvine, have unveiled a previously unknown mechanism that triggers an inflammatory immune response in cells with damaged DNA. This breakthrough not only deepens our understanding of cellular responses to DNA damage but also holds promise for developing more effective cancer treatments.
The study, published on January 6, 2025, in Nature Structural & Molecular Biology, reveals that when cells experience DNA damage due to factors like UV irradiation or certain chemotherapeutic drugs, a specific response is activated. This response prevents cells from becoming cancerous by initiating an inflammatory signal that recruits immune cells to the site of damage.
Traditionally, it was understood that when both DNA strands break, the ATM enzyme activates the protein NF-κB, leading to the production of inflammatory signals. However, this new research identifies an alternative pathway: upon DNA damage from UV exposure or specific drugs, the IRAK1 enzyme induces NF-κB to send out signals, particularly through the release of the IL-1α protein. This protein doesn’┐Įt act on the damaged cell itself but travels to neighboring cells, triggering an immune response.
The implications of this discovery are significant. By understanding how different cancer cells react to DNA damage, scientists can develop more tailored and effective therapies, potentially reducing side effects and improving patient outcomes. This research also highlights the importance of considering the immune system’┐Įs role in cancer development and treatment.
As we continue to explore the intricate mechanisms of cellular responses, such findings pave the way for innovative approaches in cancer therapy and enhance our overall understanding of human biology.

More could be read at:ATM and IRAK1 orchestrate two distinct mechanisms of NF-κB activation in response to DNA damage
Back to top
Reconstructing the Past: How Scientists Built a 3D Woolly Mammoth Genome
A groundbreaking study published in Cell on July 11, 2024, has pushed the boundaries of paleogenomics by reconstructing the 3D genome of a woolly mammoth from a 52,000-year-old fossil. This achievement marks a major step in understanding how extinct species regulated their genes and adapted to extreme environments.
Unlike previous genomic reconstructions that focused on linear DNA sequences, this study utilized Hi-C sequencing, a cutting-edge technique that maps how chromosomes fold in three-dimensional space. This approach provided a more accurate representation of how mammoth genes interacted within the nucleus, revealing key differences from their closest living relatives, Asian elephants. Notably, the researchers identified structural variations in genes related to hair growth, fat metabolism, and cold adaptation, helping explain how these Ice Age giants survived freezing conditions.
Beyond the fascination of bringing mammoths ’┐Įback to life’┐Į at the genetic level, this study has far-reaching implications for molecular biology. Understanding ancient gene regulation could shed light on modern species’┐Į evolution, climate adaptations, and even potential de-extinction efforts. Moreover, the successful retrieval of intact chromosomal structures from ancient remains raises the possibility of applying similar techniques to other extinct species, offering a new window into Earth’┐Įs evolutionary history.
This research is a testament to how rapidly genomic technology is advancing. Just a few years ago, sequencing ancient DNA was an immense challenge’┐Įnow, we’┐Įre mapping the three-dimensional organization of entire genomes from species that vanished thousands of years ago. As scientists refine these methods, we may soon uncover even more secrets hidden within the genetic blueprints of the past.
For those eager to dive into the original study, check out the full article in Cell: Three-dimensional genome architecture persists in a 52,000-year-old woolly mammoth skin sample
Back to top
Reviving Ancient Molecules: A New Frontier in Antimicrobial Research
In a groundbreaking study published in June 2024, researchers from the University of Pennsylvania have pioneered a field termed ’┐Įmolecular de-extinction,’┐Į aiming to resurrect ancient molecules from extinct species to address contemporary medical challenges. Led by bioengineer César de la Fuente, the team utilized artificial intelligence to analyze the genomes of long-extinct organisms, identifying and synthesizing proteins with potential antimicrobial properties.
Harnessing the Past to Combat Present Threats
The researchers focused on the proteomes’┐Įthe entire set of proteins expressed by a genome’┐Įof species such as woolly mammoths, ancient sea cows, and Neanderthals. By scanning these ancient genomes, they identified approximately 37,000 molecular fragments with potential antimicrobial activity. Out of these, 69 compounds were synthesized in the laboratory, with several demonstrating effectiveness against bacterial infections in preliminary tests.
Promising Results and Future Applications
Notably, peptides derived from the woolly mammoth (’┐Įmammuthusin-2’┐Į), the straight-tusked elephant (’┐Įelephasin-2’┐Į), and the giant elk (’┐Įmegalocerin-1’┐Į) exhibited significant antimicrobial efficacy. In laboratory settings, these peptides not only inhibited bacterial growth in Petri dishes but also reduced infections in mouse models. Some were found to be as effective as polymyxin B, an antibiotic of last resort.
Ethical and Legal Considerations
While the scientific community is optimistic about the therapeutic potential of these ancient molecules, the research also raises ethical and philosophical questions. The concept of resurrecting molecules from extinct species prompts discussions about human intervention in nature and our responsibilities as stewards of the biological world. Additionally, the patentability of these resurrected molecules presents a legal grey area, as they occupy a space between naturally occurring substances and synthetic compounds.
A Glimpse into the Future
This innovative approach not only offers a novel solution to the growing issue of antibiotic resistance but also exemplifies how lessons from the past can inform and enhance modern medicine. As research progresses, these ancient molecules may become integral components in the development of new antimicrobial therapies, bridging the gap between extinct biological entities and contemporary healthcare solutions.

For more detailed information, you can refer to the original publication:Deep-learning-enabled antibiotic discovery through molecular de-extinction
Back to top
In a landmark achievement, NASA’┐Įs OSIRIS-REx mission has unveiled compelling evidence about the early solar system’┐Įs role in the emergence of life on Earth. Samples retrieved from asteroid Bennu have been found to contain all five nucleobases’┐Įthe essential building blocks of DNA and RNA’┐Įalong with a variety of other organic molecules. These findings bolster the theory that asteroids may have delivered crucial ingredients for life to our planet.
The OSIRIS-REx Mission and Sample Collection
Launched in 2016, the OSIRIS-REx spacecraft embarked on a mission to study Bennu, a carbon-rich asteroid that has remained relatively unchanged since the early solar system. In October 2020, the spacecraft successfully collected approximately 121.6 grams of material from Bennu’┐Įs surface and returned it to Earth in September 2023. This mission marks the largest asteroid sample ever brought back to our planet.
Revelation of Life’┐Įs Building Blocks
Upon meticulous analysis, scientists discovered that the Bennu samples contained all five nucleobases: adenine, guanine, cytosine, thymine, and uracil. These organic compounds are fundamental to the formation of DNA and RNA, the molecules responsible for storing and transmitting genetic information in all known forms of life. The presence of these nucleobases in extraterrestrial material provides strong evidence that the basic components necessary for life are not unique to Earth and may be widespread throughout the cosmos.
Implications for the Origins of Life
The discovery of these organic molecules on Bennu supports the hypothesis that asteroids and other celestial bodies played a significant role in delivering life’┐Įs precursors to Earth. This process, known as panspermia, suggests that the early solar system was a dynamic environment where organic compounds could form and be transported across vast distances, seeding planets with the necessary ingredients for life. The findings from Bennu imply that such processes could have occurred elsewhere, increasing the possibility that life exists beyond our planet.
A Watery Past on Bennu
In addition to organic molecules, the Bennu samples revealed minerals that form in the presence of water, such as carbonates and hydrated minerals. These minerals indicate that Bennu’┐Įs parent body experienced interactions with liquid water, creating a briny environment where organic compounds could have undergone further chemical reactions. This aqueous history enhances our understanding of the conditions that may lead to the development of life-supporting molecules.
Broader Impact on Astrobiology
The OSIRIS-REx mission’┐Įs findings have profound implications for the field of astrobiology. By confirming that essential organic compounds can form and survive in space, and potentially be delivered to planetary surfaces, the study strengthens the argument that life’┐Įor at least its building blocks’┐Įcould be more common in the universe than previously thought. These insights pave the way for future missions aimed at detecting signs of life on other celestial bodies, such as Mars, Europa, and Enceladus.

For more detailed information, you can refer to the original publication:NASA’┐Įs Asteroid Bennu Sample Reveals Mix of Life’┐Įs Ingredients
Back to top
The Road to Chloroplast Editing: A New Frontier
While CRISPR-CasMito’┐Įs success with mitochondrial DNA is revolutionary, researchers are already eyeing its potential for chloroplast genomes. Chloroplasts, like mitochondria, have their own DNA and play a critical role in photosynthesis and plant metabolism. Editing chloroplast DNA could revolutionize agriculture by enhancing crop yields, drought resistance, and carbon capture efficiency. Early experiments in Arabidopsis thaliana suggest that CasMito-derived systems can target chloroplast genes, though delivery remains a hurdle due to plant cell walls. If optimized, this could lead to climate-resilient crops and bioengineered plants tailored for sustainable energy production.
Technical Challenges and Innovations
The team’┐Įs breakthrough wasn’┐Įt without obstacles. Key advancements included:
’┐Į Protein Engineering: CasMito was modified to recognize mitochondrial-specific sequences while avoiding nuclear DNA.
’┐Į Delivery Breakthroughs: Lipid nanoparticles were coated with mitochondria-targeting peptides, ensuring precise delivery.
’┐Į Minimizing Off-Target Effects: Single-molecule tracking confirmed CRISPR-CasMito’┐Įs specificity, addressing long-standing concerns about unintended edits.
These innovations not only solved mitochondrial challenges but also created a blueprint for editing other organelles.
From Lab to Clinic: What’┐Įs Ahead?
Human clinical trials for mitochondrial therapies are slated to begin in 2026, focusing on Leigh syndrome and MELAS. The FDA has granted CRISPR-CasMito ’┐ĮFast Track’┐Į designation, accelerating regulatory review. Meanwhile, biotech startups like Mitogen Therapeutics are licensing the technology to develop in vivo treatments. However, challenges persist:
’┐Į Long-Term Safety: Will edited mtDNA remain stable over decades?
’┐Į Germline Ethics: Editing embryos to eradicate mitochondrial diseases remains controversial, echoing debates around nuclear CRISPR babies.
Voices from the Scientific Community
Dr. Jennifer Doudna, CRISPR pioneer, hailed the work as ’┐Įa watershed moment for genetic medicine,’┐Į while bioethicist Dr. Henry Greely urged caution: ’┐ĮMitochondrial editing is a double-edged sword. We need global consensus on its use before clinical adoption.’┐Į
Beyond Biology: Implications for Climate and Industry
CRISPR-CasMito’┐Įs impact extends beyond healthcare:
’┐Į Bioenergy: Editing algal mitochondria could optimize lipid production for cleaner biofuels.
’┐Į Carbon Capture: Engineered plants with enhanced photosynthetic efficiency might combat climate change.
Final Thoughts
CRISPR-CasMito isn’┐Įt just rewriting mitochondrial DNA’┐Įit’┐Įs rewriting the rules of genetic engineering. As we navigate this brave new world, collaboration across disciplines will be vital to balance innovation with ethics.
Stay curious, stay informed. Explore more breakthroughs at our Blog.
The original work could be read at:Precision mitochondrial DNA editing with high-fidelity DddA-derived base editors

Back to top
A Groundbreaking Leap in Quantum Computing: February's Scientific Breakthrough
In the ever-evolving world of science and technology, February brought us a remarkable breakthrough in the field of quantum computing. Researchers at the University of Science and Technology of China, led by Professor Jian-Wei Pan, have achieved a significant milestone by demonstrating the world's first quantum computer capable of performing calculations at a speed unattainable by classical computers.
Published in the prestigious journal Physical Review Letters on February 18th, 2021, the study showcases the team's development of a 66-qubit programmable superconducting quantum processor named "Zuchongzhi." This processor has successfully demonstrated quantum supremacy by solving a specific task in just 200 seconds, a feat that would take the world's fastest supercomputer, Fugaku, approximately 8 years to accomplish.
The task in question involved sampling the output of a random quantum circuit, a problem that grows exponentially more complex as the number of qubits increases. The Zuchongzhi processor's ability to tackle this challenge with such efficiency marks a significant step forward in the quest to harness the power of quantum computing for practical applications.
Quantum computers operate on the principles of quantum mechanics, utilizing qubits that can exist in multiple states simultaneously, unlike classical bits that are limited to either 0 or 1. This unique property allows quantum computers to perform certain calculations at an exponentially faster rate than their classical counterparts, opening up new possibilities in fields such as cryptography, drug discovery, and artificial intelligence.
While the Zuchongzhi processor's achievement is a remarkable feat, it is important to note that the road to practical, large-scale quantum computing is still fraught with challenges. Quantum systems are highly sensitive to environmental interference, and maintaining the delicate quantum states required for computation remains a significant hurdle. However, the progress made by Professor Pan's team serves as a testament to the rapid advancements being made in the field and offers a glimpse into the transformative potential of quantum computing.
As we continue to push the boundaries of scientific discovery, breakthroughs like the Zuchongzhi processor remind us of the incredible potential that lies ahead. The future of quantum computing is bright, and February's groundbreaking achievement brings us one step closer to unlocking its full potential.
More could be read on the original work at: Phase-Programmable Gaussian Boson Sampling Using Stimulated Squeezed Light

Back to top
A Breakthrough in Genetic Repair: RNA That Fixes DNA Damage
Science continues to push the boundaries of what we believe is possible, and 2025 is already proving to be a monumental year for genetic research. One of the most exciting breakthroughs to emerge recently is the discovery of a unique RNA molecule capable of repairing DNA damage’┐Įa finding that could revolutionize medicine and our understanding of cellular resilience.
Researchers have identified a novel RNA-guided system that actively detects and repairs breaks in DNA strands, a process critical to preventing mutations that can lead to diseases like cancer. Unlike traditional repair mechanisms that rely heavily on proteins, this RNA molecule takes center stage, guiding the repair process with remarkable precision. Preliminary studies suggest it can mend DNA damage caused by environmental factors, such as UV radiation or chemical exposure, potentially reducing the risk of genetic disorders.
What makes this discovery a game-changer? For one, its efficiency. The RNA system appears to work faster and with fewer errors than some existing cellular repair pathways. Additionally, its mechanism is distinct from well-known tools like CRISPR, offering a complementary approach that could be less prone to off-target effects. Scientists are already speculating about its applications’┐Įimagine therapies that enhance our cells’┐Į natural ability to fix themselves, or preventative treatments that bolster DNA stability in at-risk populations.
This breakthrough builds on decades of RNA research, but it’┐Įs the first time scientists have observed an RNA molecule taking such a direct role in DNA repair. The implications are vast: from personalized medicine to anti-aging treatments, this could open doors we didn’┐Įt even know existed. While it’┐Įs still early days, with clinical trials likely years away, the discovery marks a pivotal moment in molecular biology.
At SCICC, we’┐Įre thrilled to follow these advancements and share them with you. Science isn’┐Įt just about understanding the world’┐Įit’┐Įs about reshaping it for the better. Stay tuned as we keep an eye on how this RNA breakthrough unfolds!
To read the original published work regarding this discovery you could access the following link: NEAT1 promotes genome stability via m6A methylation-dependent regulation of CHD4

Back to top
CRISPR Breakthrough: A New Tool for Editing RNA
In the dynamic landscape of genetic research, a significant advancement was made in the last six months that could transform the way we approach genetic diseases. A team of scientists from the Broad Institute of MIT and Harvard, along with collaborators from the University of California, San Diego, have developed a new CRISPR-based tool capable of editing RNA. This breakthrough, published in the journal Science in October 2017, opens up new possibilities for treating a wide range of genetic disorders.
The Innovation: REPAIR (RNA Editing for Programmable A to I Replacement)
The new tool, named REPAIR, stands for RNA Editing for Programmable A to I Replacement. Unlike traditional CRISPR-Cas9, which edits DNA, REPAIR targets RNA, offering a more flexible and potentially safer approach to genetic editing. The system uses a modified Cas13 enzyme, which, when combined with an adenosine deaminase, can convert adenine (A) to inosine (I) in RNA molecules. This change can alter the genetic code and potentially correct disease-causing mutations.
Potential Applications
The ability to edit RNA has several advantages over DNA editing. RNA editing is transient, meaning the changes do not affect the genome and are not passed on to future generations. This makes it a promising approach for treating genetic diseases without the risk of off-target effects that can occur with DNA editing. Potential applications include treating neurodegenerative diseases, such as ALS and Huntington's disease, as well as other conditions caused by RNA mis-splicing or mutations.
Challenges and Future Directions
While the development of REPAIR is a significant step forward, there are still challenges to overcome. The efficiency and specificity of RNA editing need to be improved, and the delivery of the editing machinery to target cells in the body remains a hurdle. Researchers are now working on optimizing the system and exploring its therapeutic potential in preclinical models.
The development of REPAIR represents a major breakthrough in the field of genetic editing. By targeting RNA, this new tool offers a versatile and potentially safer approach to treating genetic diseases. At SCICC, we are excited to follow the progress of this innovative technology and its potential to revolutionize medicine.
Stay tuned for more updates on this groundbreaking research and its implications for the future of genetic therapy!
To read the original published work regarding this discovery you could access the following link: RNA editing with CRISPR-Cas13

Back to top
Wiring the Mind: Scientists Unveil the Most Detailed Brain Map Yet
The human brain is often called the most complex object in the known universe, but unlocking its secrets has always been a daunting challenge. In a stunning leap forward, scientists in 2024 have given us a glimpse into that complexity by completing the most detailed map of a brain ever created’┐Įnot a human one, but that of a humble fruit fly. This breakthrough, achieved by an international team after a decade of meticulous work, could pave the way for understanding how thoughts, decisions, and memories take shape in all creatures, including us.
The fruit fly (Drosophila melanogaster), a longtime favorite of researchers due to its simple yet surprisingly relatable biology, has a brain about the size of a poppy seed. Don’┐Įt let its size fool you: this tiny organ contains nearly 140,000 neurons and millions of synaptic connections, making it a miniature marvel. The team’┐Įs achievement, published in October 2024, is a full ’┐Įconnectome’┐Į’┐Įa wiring diagram showing how every neuron links to every other. To pull this off, they sliced the fly’┐Įs brain into ultra-thin sections, thinner than a human hair, and photographed each one with a high-resolution microscope. Advanced computing then stitched the images together into a 3D model, revealing a neural network more intricate than anyone imagined.
Why does this matter? Fruit flies share many genetic and neurological traits with humans, and their brain’┐Įs basic functions’┐Įlike processing sensory input or triggering behavior’┐Įmirror ours on a smaller scale. By mapping every connection, scientists can now study how signals flow through this network, offering clues about how brains compute. Early findings suggest that some neuron clusters act like ’┐Įhubs,’┐Į directing traffic in ways that could explain decision-making or learning. For neuroscience, it’┐Įs like getting the blueprints to a city we’┐Įve only seen from the outside.
The implications are huge. This connectome is already being used to simulate brain activity on computers, a step toward cracking mysteries like how memories form or why disorders like Alzheimer’┐Įs disrupt thinking. It’┐Įs not a human brain map yet’┐Įours has 86 billion neurons, a far cry from 140,000’┐Įbut it’┐Įs a proof of concept. Researchers predict that within a decade, this work could inspire AI systems that mimic biological brains more closely or even guide treatments for neurological conditions.
This discovery isn’┐Įt just a win for science; it’┐Įs a reminder of how the smallest things can unlock the biggest questions. You can read more at: Neuronal wiring diagram of an adult brain
At SCICC, we’┐Įll be watching as this tiny fly’┐Įs brain lights the path to understanding our own. Stay tuned for more mind-blowing updates!

Back to top
Supercharged Evolution: Scientists Create an ’┐ĮEvolution Engine’┐Į for Protein Design
Imagine giving Mother Nature a fast-forward button. In August 2025, researchers at Scripps Institute unveiled exactly that: an ’┐Įevolution engine’┐Į dubbed T7-ORACLE that lets proteins evolve inside cells hundreds of thousands of times faster than usual. Instead of waiting weeks or months to fine-tune a protein in the lab, this system churns out improved protein variants in mere days. One researcher even describes it as giving evolution a ’┐Įfast-forward button’┐Į. By co-opting a viral DNA copier (from bacteriophage T7) and cranking up its error rate, the team engineered bacteria so that each cell division continuously introduces mutations in target genes at 100,000’┐Į the normal rate. In practice, this means that every 20-minute division cycle of an E. coli cell is like running a fresh round of directed evolution in real time.
Fast-Forwarding Evolution Inside Cells
Building on well-known techniques like directed evolution, the team’┐Įs new platform makes the process dramatically faster. Normally, scientists would mutate DNA, grow cells, select better proteins, then repeat ’┐Į each ’┐Įgeneration’┐Į taking a week or more. T7-ORACLE does all of that inside the cell automatically. The secret is an engineered version of the T7 bacteriophage DNA polymerase, the enzyme that copies DNA. By tweaking it to be highly error-prone, the researchers caused random mutations in a ’┐Įlab Darwin’┐Į fashion. Crucially, these mutations only affect small plasmid DNA carrying the target gene, leaving the cell’┐Įs own genome intac.In one test, the team inserted a common antibiotic-resistance gene into their system. In less than a week it evolved variants able to survive 5,000’┐Į higher antibiotic doses than the starting gene. Remarkably, many of the mutations it found mirrored those seen in real-world drug-resistant bacteria, and even some new combos that were even better. As co-author Christian Diercks explains, by the end of each cell cycle ’┐Įyou get a round [of evolution] each time the cell divides ’┐Į so it really accelerates the process’┐Į. n short, T7-ORACLE turns ordinary lab microbes into rapid evolution machines that can hone any protein in days instead of months.
Implications for Science and Medicine
This advance could reshape how we develop new drugs, enzymes, and therapies. Since T7-ORACLE is compatible with standard E. coli cultures and lab workflows, scientists can ’┐Įdrop in any gene and evolve it toward whatever function [they] need’┐Į. For example, researchers could quickly evolve high-affinity antibodies to novel cancer targets, or tailor enzymes to break down pollutants. The authors note that their platform is a breakthrough for engineering therapeutic proteins for cancer, neurodegenerative diseases, and beyond. In essence, it slashes the time and labor of protein design: what used to require many cycles of human effort can now happen at cellular speed. The breakthrough echoes the popular promise of artificial protein design but takes it into living cells, marrying synthetic biology with natural selection on steroids.
In conclusion, the creation of T7-ORACLE is a major milestone in molecular biology. By harnessing the power of evolution itself and accelerating it by orders of magnitude, scientists can now explore protein variants and solutions that were previously impractical. This ’┐Įevolution engine’┐Į opens new possibilities: from outpacing antibiotic resistance to tailoring personalized enzymes and new therapeutics, the era of turbocharged molecular design is here. As the researchers put it, combining rational design with this continuous evolution system lets us discover functional molecules ’┐Įmore efficiently than ever’┐Į. The work was published in Science in August 2025, and marks a leap forward in how we engineer biology for medicine and biotechnology.
You can read more at: An Orthogonal T7 Replisome for Continuous Hypermutation and Accelerated Evolution in E. coli

Back to top
Prime editing’┐Įs leap forward: making DNA surgery safer, smarter, and closer to the clinic
By Pedro Paulo Gattai Gomes, Ph.D.
Genome editing has already reshaped molecular biology. But until recently, precision and safety remained major hurdles for therapies that must alter patients’┐Į DNA directly. In 2025 a series of related advances ’┐Į from molecular engineering to AI-guided protein design and encouraging animal studies ’┐Į delivered a decisive push: prime editing, a versatile ’┐Įsearch-and-replace’┐Į DNA editor, is becoming far more accurate and clinically practical.
Fast-Forwarding Evolution Inside Cells
Building on well-known techniques like directed evolution, the team’┐Įs new platform makes the process dramatically faster. Normally, scientists would mutate DNA, grow cells, select better proteins, then repeat ’┐Į each ’┐Įgeneration’┐Į taking a week or more. T7-ORACLE does all of that inside the cell automatically. The secret is an engineered version of the T7 bacteriophage DNA polymerase, the enzyme that copies DNA. By tweaking it to be highly error-prone, the researchers caused random mutations in a ’┐Įlab Darwin’┐Į fashion. Crucially, these mutations only affect small plasmid DNA carrying the target gene, leaving the cell’┐Įs own genome intac.In one test, the team inserted a common antibiotic-resistance gene into their system. In less than a week it evolved variants able to survive 5,000’┐Į higher antibiotic doses than the starting gene. Remarkably, many of the mutations it found mirrored those seen in real-world drug-resistant bacteria, and even some new combos that were even better. As co-author Christian Diercks explains, by the end of each cell cycle ’┐Įyou get a round [of evolution] each time the cell divides ’┐Į so it really accelerates the process’┐Į. n short, T7-ORACLE turns ordinary lab microbes into rapid evolution machines that can hone any protein in days instead of months.
Here’┐Įs what changed, why it matters, and what to watch next.
What is prime editing ’┐Į in one paragraph
Prime editing is a CRISPR-derived approach that uses a nickase Cas9 fused to a reverse transcriptase plus a specialized guide RNA (pegRNA) to write new DNA sequences into a target site without making double-strand breaks. That design lets it install replacements, insertions, or deletions with much greater flexibility than older base editors and with fewer unintended edits than nuclease-based CRISPR. For a technical overview of the mechanism and its early developments, see recent reviews. (Li et al - 2025)
The 2025 breakthroughs ’┐Į three pillars
1) Dramatically lower error rates from protein and protocol engineering
Researchers at MIT reported a new set of modifications to the prime editor system that reduce undesired edits by orders of magnitude ’┐Į the team described reductions in error rates that make prime editing much safer for potential therapeutic use. The work combines careful enzyme engineering with improved pegRNA designs and delivery considerations; the net result is a system that introduces far fewer off-target or bystander changes than earlier prime editors. That advance directly addresses one of the biggest safety concerns for clinical genome editing.
2) AI-aided design to boost efficiency and specificity
Teams using modern protein-design and machine learning methods generated small ’┐Įbinders’┐Į and accessory proteins that improve the activity and fidelity of prime editors. A high-profile Cell paper showed that AI-designed small binders can enhance prime editing performance at difficult targets, effectively giving scientists new molecular ’┐Įtools’┐Į that steer the editing reaction in the right direction. This marriage of ML and molecular engineering accelerates optimization in cases where trial-and-error would be slow or impractical.
3) Therapeutic proof-of-concepts in animals and personalized workflows
Multiple labs reported successful in vivo applications of prime editing in disease models in 2025. Examples include rescue of retinal degeneration in mouse models and correction of neurological disease mutations in mice ’┐Į studies that show not only molecular correction but also functional improvement. Separately, clinical teams are developing workflows for patient-specific in vivo editing for ultra-rare variants, demonstrating the logistical and regulatory paths that will be needed to bring such therapies to human patients. These animal and translational studies together create a credible bridge from bench discoveries to first-in-human applications.
Why this matters?
Safety: Lower error rates reduce the risk of unintended mutations that could be harmful ’┐Į a key prerequisite for therapeutic use.
Versatility: Prime editing can install many kinds of edits (substitutions, small insertions/deletions) that are relevant across hundreds of monogenic diseases.
Speed of optimization: AI-guided design shortens the cycle between idea and usable editing reagent, enabling rapid tailoring to hard targets.
Clinical momentum: Successful animal studies and patient-specific workflows show real potential for individualized therapies for rare genetic disorders.
Caveats - what still needs work?
Delivery into the right tissues safely and efficiently remains a major bottleneck for many diseases (e.g., CNS, muscle, some internal organs).
Long-term safety and immune responses against editor proteins or delivery vehicles must be fully characterized in longer-term animal studies and clinical trials.
Regulatory and manufacturing pathways for individualized (N=1) therapies are being paved, but scaling and access are still unresolved challenges. See the recent translational reports for ongoing work in these areas.
Where the field is likely headed over the next 3’┐Į5 years
Expect an accelerating pipeline: more engineered editors with lower off-target activity, broader adoption of AI in editor design, better tissue-targeted delivery vehicles (LNPs, viral vectors with improved tropism), and an expansion of early clinical trials targeting rare but severe monogenic conditions. Regulatory frameworks are also adapting to individualized gene therapies, so translational roadmaps are becoming more concrete.
Bottom line
2025 marks a turning point for prime editing. Rather than being a promising but delicate lab tool, prime editing is rapidly maturing into a safer, more tunable technology with genuine translational traction. If delivery and long-term safety continue to improve, we may soon see a wave of clinical trials testing personalized genome corrections for diseases that were once untreatable.
References
1.MIT / ScienceDaily ’┐Į ’┐ĮMIT scientists have found a way to make gene editing far safer and more accurate’┐Į (2025).’┐ĮMIT’┐Įs new precision gene editing tool could transform medicine
2.SciTechDaily ’┐Į ’┐ĮMIT scientists unlock a new level of precision in gene editing.’┐Į (Oct 2025).MIT Scientists Unlock a New Level of Precision in Gene Editing
3.Park J-C. et al., Cell ’┐Į AI-generated binder improves prime editing efficiency and specificity (2025).AI-generated MLH1 small binder improves prime editing efficiency
4.Fu Y. et al., Nature Communications ’┐Į In vivo prime editing rescues photoreceptor degeneration in mice (2025).In vivo prime editing rescues photoreceptor degeneration in nonsense mutant retinitis pigmentosa
5.Musunuru K. et al., NEJM ’┐Į Patient-specific in vivo gene editing workflow for ultrarare variants (2025).Patient-Specific In Vivo Gene Editing to Treat a Rare Genetic Disease
6.Review: Prime editing overviews and mechanism (2025) ’┐Į Li M. et al., PMCID article.Prime Editing: A Revolutionary Technology for Precise Treatment of Genetic Disorders

Back to top
A Denisovan in the Taiwan Strait: how a jawbone is reshaping the map of human evolution
By Pedro Paulo Gattai Gomes, Ph.D.
Sometimes a single fossil can change the boundaries of what we think we know. That’┐Įs exactly what happened in 2025 when researchers announced that a mysterious jawbone’┐Įknown as Penghu 1, recovered from the seafloor of the Penghu Channel off Taiwan’┐Įbelongs to a Denisovan individual. The claim rests not on DNA (which the warm, humid conditions destroyed) but on palaeoproteomics’┐Įthe analysis of ancient proteins preserved in tooth enamel and bone. This identification pushes the confirmed geographic range of Denisovans thousands of kilometres farther south and shows they occupied a far wider variety of environments than previously documented.
What was found and how it was identified
Penghu 1 was dredged from seabed sediments before 2008, entered the antiquities market, and ultimately made its way to Taiwan’┐Įs National Museum of Natural Science. Attempts to recover ancient DNA failed (not surprising in subtropical contexts), so scientists turned to proteins’┐Įbiomolecules that can survive far longer than DNA in some fossil contexts. Using targeted palaeoproteomic sequencing of dental enamel and other tissues, the team detected protein variants that match those found in other Denisovan material, allowing a confident taxonomic assignment even in the absence of recoverable DNA. That methodological pivot’┐Įproteins as a molecular archive’┐Įwas decisive in this study. .
Why this matters for human evolution
Before this work, securely identified Denisovan remains were extremely sparse and geographically clustered (Denisova Cave in Siberia and the Xiahe mandible on the Tibetan Plateau being the best-known examples). Confirming Penghu 1 as Denisovan demonstrates that Denisovans were not restricted to high-altitude or cold environments; they also occupied low-latitude, warm coastal regions in East Asia. That breadth implies ecological flexibility and raises new questions about Denisovan population structure, migration routes, and interactions with contemporaneous humans (Homo sapiens) and Neanderthals. The find therefore widens both the map and the set of hypotheses we can test about archaic human diversity in Asia.
Broader scientific implications
1) Proteins as a tool
This study is a clear demonstration that palaeoproteomics can identify species where DNA cannot be recovered, making previously ambiguous fossils testable with molecular methods. That opens many new possibilities in regions (tropical/subtropical) and time intervals where DNA preservation is poor.
2) Geographic and ecological range
The discovery supports a model in which Denisovans occupied diverse habitats across eastern Eurasia, so genomic signals of Denisovan ancestry in present-day populations (e.g., in Melanesia and parts of South and Southeast Asia) may reflect a complex history of multiple Denisovan groups and admixture events.
3) Reassessing fragmentary fossils
Many isolated or puzzling fossils in museum collections could yield new answers if reanalyzed with palaeoproteomics, potentially rewriting regional hominin histories without new excavations
Why this matters?
Safety: Lower error rates reduce the risk of unintended mutations that could be harmful ’┐Į a key prerequisite for therapeutic use.
Versatility: Prime editing can install many kinds of edits (substitutions, small insertions/deletions) that are relevant across hundreds of monogenic diseases.
Speed of optimization: AI-guided design shortens the cycle between idea and usable editing reagent, enabling rapid tailoring to hard targets.
Clinical momentum: Successful animal studies and patient-specific workflows show real potential for individualized therapies for rare genetic disorders.
Caveats and next steps
Dating uncertainty: Because Penghu 1 was recovered from the seafloor (not a stratified excavation), pinning an exact age remains challenging; estimates vary depending on sea-level and geological models. Better contextual dating would sharpen interpretations of when Denisovans reached the region.
Sample size and diversity: Denisovan material is still rare. One jawbone does not capture their full variation’┐Įmore finds and more proteomic/genomic data are needed to map Denisovan population structure and chronology across Asia.
Interdisciplinary follow-up: Integrating palaeoproteomics with archaeology, paleoenvironmental reconstructions, and ancient-DNA surveys of sediments and associated fauna will help reconstruct the lifeways, diets, and movements of Denisovan groups.
Bottom line
Penghu 1 is a striking reminder that museum drawers, antique shops, and the sea bed can still yield discoveries that reshape big-picture questions. By confirming a Denisovan identity through protein analysis, researchers have extended the known reach of these enigmatic archaic humans into subtropical East Asia’┐Įand, equally important, showcased a molecular toolkit that will let us test many more old mysteries in regions where DNA cannot survive.
References
1.Tsutaya T., Sawafuji R., Welker F., et al. A male Denisovan mandible from Pleistocene Taiwan. Science. 2025’┐ĮA male Denisovan mandible from Pleistocene Taiwan
2.Nature (News & Features). Who were the ancient Denisovans? Fossils reveal secrets. May 2025.Who were the ancient Denisovans? Fossils reveal secrets about the mysterious humans
3.Natural History Museum (UK). Denisovan jawbone helps to reveal appearance of ancient human species. Apr 2025.Denisovan jawbone helps to reveal appearance of ancient human species
Back to top
Vascularized organoids: the lab-grown tissues that might finally cross the clinicÆs threshold
By Pedro Paulo Gattai Gomes, Ph.D.
Imagine a miniature human heart, lung or brain Ś grown from stem cells in a dish Ś that already has blood vessels, behaves more like real tissue, and can be joined to a living animal so researchers can watch it develop and function. That future just moved markedly closer to the present. Over the last year researchers have published a string of studies and perspectives showing that organoids (3-D tissue models grown from pluripotent stem cells) can be engineered or induced to form functional vasculature, integrate with host circulation after transplantation, and reach levels of maturity that make them far more useful for disease modelling and therapeutic research than ever before.
What changed Ś and why it matters
Organoids have been around for over a decade as powerful models of human development and disease. Until recently, a major bottleneck was their lack of a vascular (blood-vessel) network: without vessels, organoids are small, metabolically stressed and immature, limiting how faithfully they reproduce adult tissue physiology or long-term disease processes. Engineering reliable vasculature Ś either by coaxing endothelial cells to self-assemble inside organoids, by microfluidic perfusion, or by transplanting organoids into a living host so the hostÆs blood vessels grow into them Ś removes that bottleneck and unlocks far richer biology.
The recent advances
A major, coordinated perspective led by Pașca and colleagues lays out a field-wide framework for neural organoids, assembloids and transplantation studies Ś including standardization, quality control and ethical considerations Ś reflecting how close some teams now are to translational applications. At the experimental front, several groups have demonstrated methods that reliably generate vascularized organoids or achieve host-mediated vascularization after transplantation, and have published protocols for assessing functional integration (electrophysiology, perfusion, host circuit engagement) in animal models. These papers together move organoids from isolated micro-models toward vascularized tissues that can be interrogated in vivo.
How researchers build vessels inside organoids
There are a few complementary technical routes that researchers are using:
- Co-differentiation or co-culture: adding endothelial progenitors during organoid formation so vessel networks form as the tissue self-organizes.
- Bioengineering and perfusion: using microfluidic scaffolds or ōkirigamiö electronics to provide sustained perfusion and chronic electrophysiological access.
- Transplantation and host vascularization: implanting organoids into immunocompromised rodents (or other permissive hosts) so the hostÆs vasculature invades and perfuses the graft, dramatically increasing survival and maturation.
Each approach has strengths: engineered vasculature is controllable in vitro; transplantation yields rapid, physiologic perfusion and host-cell interactions. Combining methods and rigorous reporting (the Pașca framework stresses exactly this) is whatÆs accelerating reproducibility and impact.
Why this is truly groundbreaking
Vascularized organoids change the calculus for several high-impact applications:
- Disease modelling: Organoids that mimic adult tissue physiology can model complex chronic diseases (neurodegeneration, fibrotic lung disease, cardiomyopathies) with higher fidelity.
- Drug discovery and toxicology: Perfused organoids permit realistic pharmacokinetics and multi-cell interactions, improving preclinical prediction.
- Regenerative medicine (long-term potential): If safety, scale-up, and immune-compatibility hurdles are solved, vascularized organoids could serve as building blocks for tissue replacement or for enhancing regenerative protocols.
Caveats and the road ahead
Important barriers remain. Immune rejection, the risk of unwanted proliferation, long-term functional stability, vascular architecture matching native organs, and manufacturing at clinical scale are nontrivial challenges. Moreover, the ethical and regulatory landscape needs harmonized standards for reporting, consenting, and transplantation Ś exactly the gaps the Pașca et al. paper is calling the field to address. Careful, standardized preclinical work and transparent reporting will be essential before any clinical translation.
Bottom line
The field is no longer talking only about ōmini-organs in a dish.ö ItÆs engineering tissues that include lifelike blood networks and that can meaningfully interact with living hosts Ś a shift that elevates organoids from powerful laboratory curiosities to legitimate translational tools. That shift doesnÆt guarantee immediate cures, but it makes several previously speculative applications (better human-specific disease models, more predictive drug testing, and eventual regenerative therapies) substantially more plausible in the coming decade.
References
1.Pașca SP, et al. A framework for neural organoids, assembloids and transplantation studies. Nature, 2025.A framework for neural organoids, assembloids and transplantation studies
2.Kistemaker L, et al. Vascularized human brain organoids: current possibilities and prospects. Trends Biotechnol, 2025.Vascularized human brain organoids: current possibilities and prospects
3.Pașca Labs. Key publications. Stanford Medicine, 2025.Key publications
4.Kelley, K. W. et al. Host circuit engagement of human cortical organoids transplanted in rodents. Nature Protocols, 2024. Host circuit engagement of human cortical organoids transplanted in rodents
5.Mallapaty, S. Mini hearts, lungs and livers made in lab now grow their own blood vessels. Nature (News), 2025. Mini hearts, lungs and livers made in lab now grow their own blood vessels.
Back to top
Rewinding the epigenetic clock: partial and chemical reprogramming show real rejuvenation in cells and animals
By Pedro Paulo Gattai Gomes, Ph.D.
What if we could roll back some hallmarks of cellular aging without turning cells into stem cells? Over the past few years Ś and with several notable advances in 2024¢2025 Ś researchers have shown that partial reprogramming (brief, controlled activation of Yamanaka-like factors) and chemical reprogramming (small-molecule cocktails that induce a youthful epigenetic state) can reverse molecular signs of aging in cells, restore tissue function in disease models, and in some cases improve lifespan and healthspan in mice. These experiments suggest a new, practical route to epigenetic rejuvenation that aims to restore youthful function while avoiding full dedifferentiation and the tumor risks of classic reprogramming.
What the breakthrough actually did
Partial reprogramming uses transient or cyclic expression of pluripotency-associated transcription factors (for example variants of Oct4, Sox2, Klf4 with or without c-Myc) for short pulses instead of continuous expression. That limited exposure can reset multiple aging markers Ś epigenetic clocks, transcriptomes, mitochondrial readouts, chromatin marks, and senescence-associated programs Ś while preserving cell identity. In several preclinical reports, cyclic OSKM/OSK regimens improved tissue repair, lowered inflammatory signatures, and reduced molecular measures of biological age; in targeted studies, they even improved cognitive function and extended lifespan in specific mouse paradigms.
Parallel to genetic approaches, chemical reprogramming uses stepwise cocktails of small molecules to induce a plastic, rejuvenated state without delivering exogenous transcription factor genes. Recent studies report that carefully designed small-molecule cocktails can rejuvenate human cells in vitro at the multi-omic level and extend lifespan in simple animal models, offering a non-genetic route to epigenetic reset that is inherently more compatible with systemic delivery.
Why this matters (practical implications)
A reversible, targeted path to rejuvenation. Instead of attempting to ōfixö every genomic lesion, partial reprogramming targets the epigenome and transcriptional networks that underlie many aging phenotypes Ś and in animal models this produces measurable functional improvements (better tissue repair, cognition, metabolic markers). That makes partial reprogramming an attractive way to treat age-related organ decline and certain degenerative diseases.
Non-integrative and chemical routes ease translation.Chemical reprogramming avoids gene therapy vectors altogether; transient mRNA or AAV strategies for controlled OSK pulses reduce long-term exposure to reprogramming factors Ś both features that improve clinical feasibility.
Biomarker-driven development.Advances in multi-omic aging clocks and functional assays now let researchers quantify rejuvenation effects robustly across tissues Ś a prerequisite for rationally moving these interventions toward human trials.
Important caveats and safety concerns
This field balances promise with real risks. Continuous or poorly controlled reprogramming can cause loss of cell identity, uncontrolled proliferation, and teratoma formation. Even with cyclic or partial regimens, effects vary by tissue, sex, and age, and the long-term consequences (including rare oncogenic events or genomic instability) need rigorous, long-duration study. Delivery remains a major translational hurdle Ś getting precise, intermittent expression or the right small-molecule concentrations to the intended tissues without off-target effects is nontrivial. For these reasons, most work so far is preclinical and cautious.
Where the field is headed
Expect three simultaneous thrusts over the next few years: (1) mechanistic dissection to separate rejuvenation pathways from dedifferentiation programs (so we can keep benefits and lose the risks), (2) safer delivery technologies (transient mRNA, tissue-targeted AAVs, or systemic small-molecule regimens), and (3) rigorous, biomarker-driven preclinical studies to map durability and rare adverse events. If those succeed, targeted trials for organ-specific indications (for example, retinal or cardiac repair, or age-related cognitive decline) will likely precede any attempt at whole-body rejuvenation.
References
1.Y³cel A.D., et al. The long and winding road of reprogramming-induced rejuvenation. Nature, 2024.The long and winding road of reprogramming-induced rejuvenation
2.Sahu S.K., et al. Targeted partial reprogramming of age-associated cell states improves tissue function. Science Translational Medicine, 2024.Targeted partial reprogramming of age-associated cell states improves markers of health in mouse models of aging
3.Schoenfeldt L., et al. Chemical reprogramming ameliorates cellular hallmarks of aging. EMBO / Nature-family article, 2025.Chemical reprogramming ameliorates cellular hallmarks of aging and extends lifespan
4.Ivanova J. Partial reprogramming and rejuvenation: multi-omic views and translational prospects. International Journal of Molecular Sciences, 2024. Partial Reprogramming Exerts a Rejuvenating Effect on Human Mesenchymal Stem Cells That Underwent Replicative Senescence in Culture

Back to top
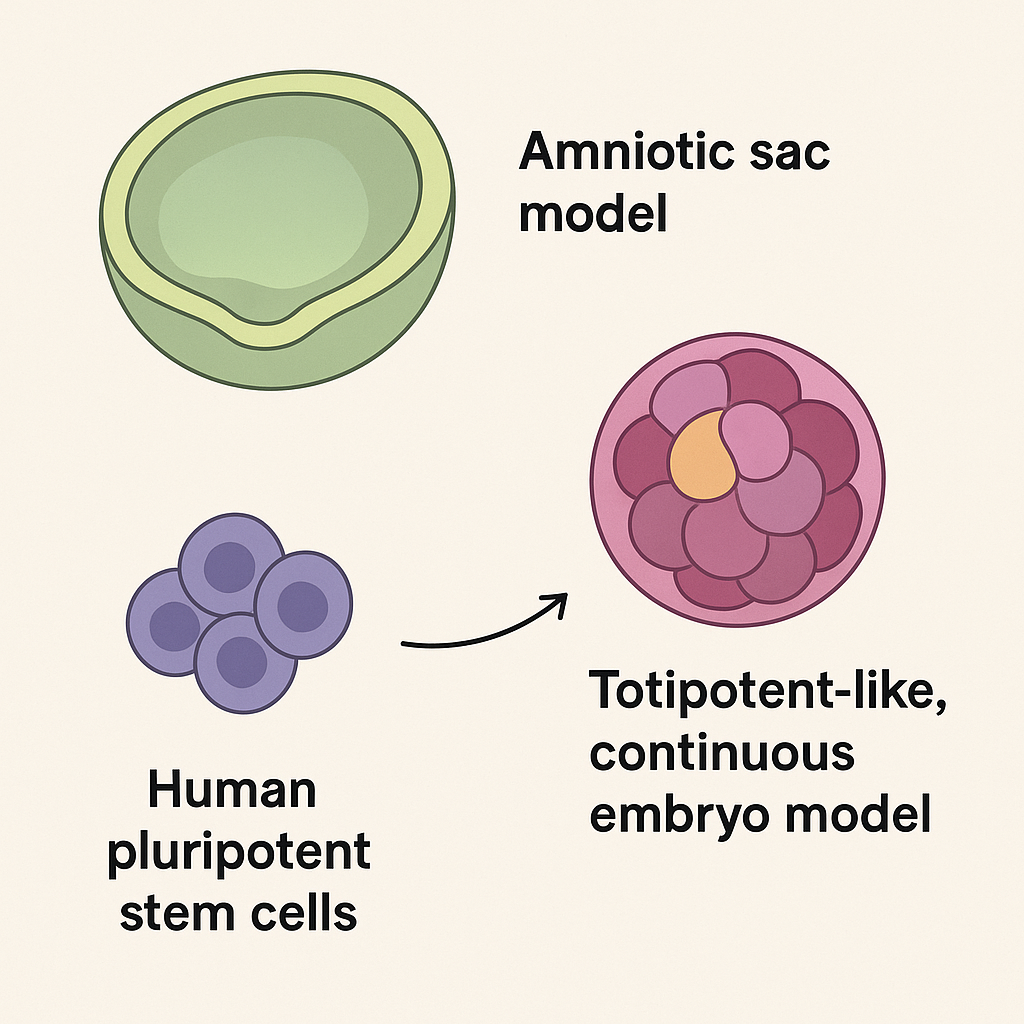
How lab-grown ōmini amniotic sacsö and next-gen embryo models are rewriting early human development
By Pedro Paulo Gattai Gomes, Ph.D.
Researchers this year reported striking advances in stem-cell¢derived embryo models: teams coaxed human stem cells to self-assemble into more advanced amniotic-sac structures and described new totipotent-like cell systems that push embryo-model development further along the roadmap from fertilization to organogenesis. These are not embryos made from egg + sperm, but powerful Ś and ethically sensitive Ś in-vitro models that let scientists watch the earliest stages of human development with a level of detail previously impossible.
What exactly was achieved?
Two classes of advances stand out. First, groups produced amniotic-sac models: hollow, fluid-filled structures made from human pluripotent stem cells that reproduce the physical environment surrounding a very early embryo, enabling study of how the amnion forms and protects the embryo in the first weeks. Second, researchers described totipotent-like, continuous embryo models built from carefully conditioned stem cells that more faithfully recapitulate the developmental trajectory from genome activation through gastrulation-like events Ś the crucial stage when body axes and primary tissue layers are established. Together, these models capture both the structural (extraembryonic sacs) and cell-lineage (embryonic/totipotent trajectories) aspects of early development.
Why this matters
Studying human embryos in vivo is ethically and practically limited Ś so stem-cell-based embryo models are a major new tool. They let researchers: (1) dissect the molecular signals that steer the first cell-fate decisions, (2) model causes of early pregnancy loss or congenital defects, and (3) screen drugs or environmental exposures that might affect implantation and early organogenesis. Because some models now generate extraembryonic structures (like amniotic sacs) and totipotent-like lineages, experiments can probe embryo¢maternal interface biology and early cell interactions that were previously inaccessible.
How the work was done
Teams used advanced stem-cell culture conditions that push cells into naive or totipotent-like states, combined them in defined 3-D conditions, and allowed the cells to self-organize. Careful imaging, single-cell sequencing and functional tests showed that the resulting structures express the gene programs and form the tissue-like arrangements expected in the corresponding embryonic stages. Iterative optimization Ś changing growth factors, timing, and mechanical cues Ś was essential to coax predictable, reproducible outcomes.
Caveats and the ethics question
These models are powerful but not the same as a naturally conceived embryo Ś and that distinction matters ethically, legally and scientifically. The field is rapidly evolving, and ethicists, funders and regulators are already debating new guardrails for how far and for how long such models should be developed in vitro. Transparent reporting, independent oversight, and international dialogue will be essential as the models gain physiological realism.
References
1.Nature News: ōStem cells coaxed into most advanced amniotic sacs ever.öStem cells coaxed into most advanced amniotic sacs ever grown in the lab
2.Ren Y. et al., Nature Cell Biology (2025): ōA continuous totipotent-like cell-based embryo modelö (advances in totipotent-like stem cell embryo models).A continuous totipotent-like cell-based embryo model recapitulates mouse embryogenesis from zygotic genome activation to gastrulation
3.Rosner M. et al., Stem Cell Research (2025) Ś review on stem cell¢based human embryo models and applications.Stem cell-based human embryo models: current knowledge and open questions
4.Lewis J., Journal of Medical Ethics (2025) Ś perspectives on ethical and regulatory implications of synthetic embryo research. Manipulating embryogenesis and testing forpotential: two real problems for the regulation of stem cell-based embryo models
Back to top
A new chapter for curing diabetes: CRISPR-edited cells make insulin without triggering the immune system
By Pedro Paulo Gattai Gomes, Ph.D.
Imagine a future where people with type 1 diabetes Ś or other severe forms of the disease Ś no longer depend on daily insulin injections or medications that suppress the immune system. In 2025, science moved a clear step toward that future: for the first time, CRISPR-edited cells were implanted in a human, produced insulin, and at the same time remained ōinvisibleö to the immune system.
Why this discovery matters
The major hurdle for cell therapies in diabetes Ś especially autoimmune forms Ś has been immune destruction of transplanted insulin-producing cells. With genetic editing, those cells were able to ōtrickö the immune system and avoid rejection without the need for systemic immunosuppression.
- The major hurdle for cell therapies in diabetes Ś especially autoimmune forms Ś has been immune destruction of transplanted insulin-producing cells. With genetic editing, those cells were able to ōtrickö the immune system and avoid rejection without the need for systemic immunosuppression.
- This strategy shows it may be possible to restore lost function durably and naturally Ś not just supplying exogenous insulin, but rebuilding the bodyÆs own regulated insulin production.
How it works, in broad terms
- Scientists used CRISPR to modify cells so they both express insulin and carry markers that ōcamouflageö immune activation signals. When implanted, these cells behave as if they are part of the host tissue.
- The technique is still experimental and requires rigorous safety testing and long-term follow-up. But a successful case suggests the approach is moving beyond theory toward clinical reality.
Broader implications
- Less reliance on immunosuppressants Ś Reducing or eliminating the need for drugs that dampen the immune system could lower serious side effects and long-term complications.
- Potential application to other autoimmune diseasesŚ If transplanted cells can be made immune-tolerant, the method could extend to conditions where the immune system attacks healthy tissues.
- A new therapeutic paradigmŚ Instead of treating symptoms (for example, with external insulin), the approach aims to restore natural function from within.
Challenges and what remains to be done
- Safety and long-term monitoring Ś Any genetically modified implanted cell must be followed closely for stability, immune behaviour, and potential adverse effects.
- Scalability and costŚ Producing edited cells at clinical grade and ensuring immune tolerance is complex and expensive; making this widely available will require major infrastructure investments.
- Regulation and ethicsŚ Gene-based therapies raise regulatory questions and issues of access: who will receive them, and how can safety and equity be ensured?
Conclusions
The recent demonstration that CRISPR-edited cells can produce insulin in a person without triggering immune detection is among the most promising advances of 2025 Ś a glimpse of a future where diabetes might be treated, or even cured, more definitively. If upcoming studies confirm safety and effectiveness, this could be a real turning point for regenerative medicine and for treating autoimmune diseases more broadly.
This is science doing what itÆs meant to do: restoring peopleÆs capacity to live with greater autonomy.
Reference
1.Nature News: ōHope for diabetes: CRISPR-edited cells pump out insulin in a person Ś and evade immune detection.öHope for diabetes: CRISPR-edited cells pump out insulin in a person Ś and evade immune detection
Back to top
Prime editing reaches the clinic: a quiet revolution in genetic medicine
Imagine a ōsearch-and-replaceö tool for DNA that can fix a single missing letter without blowing a hole in the genome. ThatÆs prime editing in a nutshell Ś and for the first time, it has been used successfully in people. In December 2025 the New England Journal of Medicine published first-in-human clinical data showing that an autologous stem-cell therapy using prime editing corrected the common ΔGT mutation that causes p47^phox-deficient chronic granulomatous disease (p47-CGD), restored neutrophil oxidase function, and produced early clinical benefit in treated patients.
Why this matters
Traditional CRISPR-Cas9 editing makes double-strand breaks in DNA, which can be efficient but also risky: unwanted insertions/deletions and activation of DNA damage responses are real concerns. Prime editing was invented to avoid those breaks: a Cas9 nickase is fused to a reverse transcriptase and guided by a special pegRNA that both targets and encodes the exact correction to be written into the genome. That makes it a more precise, versatile ōsearch-and-replaceö system capable of small insertions, deletions, and all 12 base-to-base conversions Ś theoretically reducing collateral damage and broadening the kinds of mutations we can treat.
What the NEJM paper showed
The NEJM report describes early clinical results from PM359 Ś Prime MedicineÆs investigational autologous CD34⁺ hematopoietic stem-cell product Ś in patients with p47-CGD. In short:
- Ex vivo prime editing of patient HSCs corrected the disease-causing mutation and the edited cells engrafted after reinfusion.
- Functional assays showed rapid restoration of NADPH oxidase activity (measured by DHR assays), reaching levels expected to provide clinical benefit within weeks.
- No immediate safety signals attributable to the editing approach were reported in the treated patients in these early data, although longer follow-up and more patients are needed to assess durability and rare events These outcomes provide the first clinical proof that prime editing can correct a disease-causing mutation in human stem cells and produce biological and early clinical effects. ThatÆs why many in the field call this a milestone Ś not the end of the road, but a major step from bench to bedside.
- Noise and scale. Maintaining coherence across hundreds (and later thousands) of qubits remains a technical challenge. Error-mitigation protocols and fine-grained calibration are essential.
- Generality. Not every useful problem will exhibit the same acceleration; mapping which real-world applications (chemistry, materials, certain optimizations) benefit most from the technique is required.
- Independent reproducibility. The community normally requires replication on different hardware and by independent groups before a result is accepted as a turning point. The authorsÆ emphasis on ōverifiabilityö is precisely a response to that demand.
- Proof of concept for ultra-rare disease treatment paradigms. This case provides a template: when a child faces imminent morbidity or mortality and a well-defined single-nucleotide cause exists, a bespoke editing intervention can be ethically and technically justified and feasibly delivered. That changes the calculus for clinicians and regulators who manage rare-disease care.
- Acceleration of regulatory thinking. Regulators and institutional review boards will need to evolve frameworks that allow expedited review of patient-specific interventions while preserving robust safety oversight and data collection requirements. The case will inform standards for CMC (chemistry, manufacturing and controls), potency assays, and post-treatment registries.
- Manufacturing and cost challenges. Single-patient GMP manufacturing remains resource-intensive. For these therapies to serve more patients, the community must invest in modular, flexible manufacturing platforms and validated, standardized preclinical packages that regulators accept as sufficient for safety screening.
- Ethical and equity considerations. Personalized editing raises thorny questions about access (who can get custom therapies), prioritization, and consent, especially in neonates. The field must develop policies to prevent inequitable access to life-saving bespoke therapies.
- Generality. A single, well-executed patient case does not prove broad safety or efficacy across genotypes, ages, or organ systems. Extrapolating from one neonate to wider populations would be scientifically unsound.
- Off-target and mosaicism concerns. Even base editors can produce off-target edits at DNA or RNA level; low-frequency events might be clinically silent initially but consequential later. Deep molecular surveillance (including unbiased genome-wide assays) is essential.
- Durability and repeatability. LNP delivery is transient; whether a single dosing episode provides durable correction for diseases with lifelong pathology must be evaluated case by case. For hepatocyte-driven metabolic disorders, hepatocyte turnover and selective advantage/disadvantage of edited cells are relevant variables.
- Efficacy in human-relevant models. Mice (even human-xenograft models) have tumor microenvironments that differ from humans. Validation in humanized stroma models and, before that, robust toxicology in large animals are needed.
- Thermal control and safety. FUS requires precise dosing; small temperature overshoots can damage healthy tissue. Protocols, sensing systems, and contingency plans must be validated under regulatory standards.
- Scale and logistics. Combining cell therapy with specialized focused-ultrasound equipment increases cost and operational complexity. Tertiary centers might implement it, but broad deployment outside referral centers will be challenging.
Strengths, caveats, and what to watch next
Strengths: the approach is targeted, precise, and uses patientsÆ own edited stem cells (which minimizes immune rejection). The early engraftment and functional correction are exactly the signals you want to see in a proof-of-concept trial.
Caveats: early-phase studies enroll few patients and follow-up is limited. Key open questions include long-term durability of the correction, low-frequency off-target edits that might escape early detection, and manufacturing scalability and cost for broader use. Independent replication, larger cohorts, and multi-center data will be essential.
What to watch: follow-up reports with longer safety data, results from additional indications (some groups are pursuing other monogenic disorders), and improvements in delivery/manufacturing that could make ex vivo prime editing faster and cheaper. Meanwhile, regulators and ethicists will closely monitor off-target analyses and long-term surveillance plans.
Why clinicians and the public should care
If prime editing continues to show safety and efficacy, it could widen the range of genetic diseases we can treat precisely: disorders caused by small deletions or point mutations are prime candidates. Because it doesnÆt rely on donor DNA templates or double-strand breaks, prime editing could offer cleaner fixes for many monogenic disorders Ś potentially converting once-lifelong conditions into curable ones with a single therapeutic intervention. ThatÆs the long-term promise; for now, the NEJM data bring that promise into the clinic for the first time.
The NEJM report is a pivotal clinical milestone: prime editing has moved from concept and preclinical excitement into the first human proof-of-principle. It doesnÆt mean every genetic disease is instantly curable, but it opens a powerful, more precise set of tools for gene therapy. Researchers, clinicians, regulators, and patients will be watching the next waves of trials closely Ś and with good reason.
References
1. Gori JL, Haddad E, Frangoul H, et al.Prime Editing for p47phox-Deficient Chronic Granulomatous Disease.N Engl J Med. 2025. DOI:10.1056/NEJMoa2509807Prime Editing for p47phox-Deficient Chronic Granulomatous Disease
2. Prime Medicine Ś press releaseannouncing NEJM publication and PM359 clinical data.Prime Medicine Announces The New England Journal of Medicine Publication of PM359 Clinical Data for the Treatment of Chronic Granulomatous Disease
3 Ledford H.World first: ultra-powerful CRISPR treatment trialled in a person.Nature (news). 2025. Ś commentary and context on the first prime-editing clinical use.World first: ultra-powerful CRISPR treatment trialled in a person
4. Broad Institute / Advanced reportssummarizing prime/base editing clinical landscape and next steps in translation.Single prime editing system could potentially treat multiple genetic diseases
5. Inside Precision Medicine and other industry coverage summarizing trial design and implications for development pathways.Prime Editing Clinical Results Published for the First Time Ever

Back to top
Molecular maps inside the cell: how cryo-ET and AI are drawing the living ōmolecular anatomyö
Imagine stepping into a cell and seeing, in 3D and at molecular resolution, where every protein complex is located Ś not an artistÆs illustration, but an experimental reconstruction of the cell itself. Recently, a set of technical and computational advances has made this idea practically achievable: cryo-electron tomography (cryo-ET) at scale, combined with artificial intelligence tools for reconstruction and automated identification, is enabling the creation of true in situ molecular atlases Ś a leap comparable to mapping the genome, but for the molecular architecture of cells and tissues.
What was demonstrated Ś a quick summary
Recent studies have shown it is possible to obtain high-resolution structures of molecular machines directly in vitrified tissue or cells, scale acquisition and processing of those volumes with automated pipelines, and Ś crucially Ś use machine-learning models to reconstruct and identify components with much less manual intervention. This opens the door to seeing biological systems Ś mitochondria, respiratory complexes, synapses, viral components inside infected cells Ś with a level of detail and spatial context that was previously almost impossible.
Why this really matters
Native context: we can now visualize proteins and complexes in their natural environment Ś membranes, cytoskeleton, and surrounding molecules Ś answering questions that isolated-structure methods (crystallography, single-particle cryo-EM) cannot.
From observation to quantitation: with increasing throughput and automation, this is no longer just ōhereÆs a beautiful volumeö; itÆs possible to quantify differences between cells, physiological states, and drug responses.
Intersection with AI: models trained on millions of particles and maps enable automatic building of atomic models and even suggest protein identities Ś cutting weeks of human work down to hours or minutes. That democratizes the method and accelerates discovery.
Key evidence (recommended reading)
In situ studies reporting high-resolution structures of respiratory complexes and other assemblies inside cells and tissues, demonstrating that functional details can be visualized in native context.
Papers describing high-throughput cryo-ET pipelines that combine FIB-milling sample preparation, automated tilt-series collection, and large-scale processing Ś prerequisites for building population-level cellular atlases.
Research and tools from the machine-learning side (for example, foundation models for cryo-EM image processing and automatic modeling tools like ModelAngelo) that automate map interpretation and atomic model building. Without this AI layer, the scale achieved today would be impractical.
Remaining limitations and challenges
Preparing tissue for cryo-ET is zero-margin work: samples need to be milled down with a focused ion beam, a fiddly, high-precision step that can itself introduce artifacts Ś and some tissue types are simply harder to make thin and pristine than others. In short, not every sample is equally cooperative.
Catching proteins that are present in vanishingly small amounts is like trying to make out a whisper across a bustling market Ś the hint of a signal is there, but easy to miss. AI tools can amplify that faint voice, yet they usually rely on huge reference libraries to guess what it might be, and those educated guesses still have to be double-checked the old-fashioned way at the bench.
And itÆs not only a technical challenge Ś itÆs an investment. Those giant microscopes, liquid-nitrogen cryo rigs and the powerful server clusters that process the data donÆt come cheap Ś and they need specialist staff to run. So instead of every lab buying and hosting the whole setup, wider access will likely come through shared core facilities, regional collaboration centers, or pay-per-use service models.
What this changes for research and medicine
This convergence of advances turns formerly speculative questions into testable hypotheses: how does the spatial organization of protein complexes change in neurodegeneration? Where, exactly, do drugs Ś or invading pathogens Ś come face-to-face with organelles inside human tissue? If we can reliably map those encounters across many cells, we could dramatically speed up the hunt for drug targets, reveal spatial biomarkers, and uncover cellular mechanisms that have been hiding in plain sight.
Conclusion Ś cautious optimism
WeÆre at an inflection point: the combination of advanced instrumentation (cryo-ET/FIB) and artificial intelligence for processing and modeling is converting high-resolution images into functional molecular maps. Technical and practical barriers remain, but the road to molecular atlases of human tissues is shorter than many expected Ś and once those atlases exist, they will transform both basic biology and translational medicine.
References
1. Zheng W, Chai P, Zhu J, Zhang K.High-resolution in situ structures of mammalian respiratory supercomplexes.Nature. 2024. DOI:10.1038/s41586-024-07488-9High-resolution in situ structures of mammalian respiratory supercomplexes
2. Jamali K, Kõll L, Zhang R, Brown A, Kimanius D, Scheres SHW. Nature. 2024. Automated model building and protein identification in cryo-EM maps. Automated model building and protein identification in cryo-EM maps
3. Heydari S, Liu J.Curr Opin Struct Biol. 2025. High-throughput cryo-electron tomography enables multiscale visualization of the inner life of microbes.High-throughput cryo-electron tomography enables multiscale visualization of the inner life of microbes
4. Majumder P, Zhang P.In situ cryo-electron microscopy and tomography of cellular and organismal samples.Curr Opin Struct Biol.2025.In situ cryo-electron microscopy and tomography of cellular and organismal samples

Back to top
Did dark matter and neutrinos just reveal a hidden handshake?
A fresh analysis that surfaced in January 2026 proposes something bold: dark matter and neutrinos Ś two of the UniverseÆs most mysterious ingredients Ś may weakly interact with each other, and that tiny interaction could explain a stubborn mismatch between early- and late-Universe measurements of how clumpy matter is (the so-called S₈ tension). If confirmed, this would force a rethink of the standard cosmological model and open an unexpected window into dark-sector physics.
What exactly was found
A team of cosmologists fit cosmological models that allow a small coupling between dark matter (DM) and neutrinos to current data sets Ś including measurements of the cosmic microwave background and recent cosmic-shear (weak gravitational lensing) surveys. They found that a nonzero DM¢neutrino interaction strength improves the fit to the data and can alleviate the S₈ tension: the late-time Universe looks slightly less clumped than early-time probes predict, and a subtle DM¢neutrino drag during structure formation can naturally produce that effect. The idea and evidence were laid out in a recent preprint and covered in the press in early January 2026.
Why this could be a game-changer
It changes what ōdark matterö could be. For decades weÆve modeled dark matter as practically collisionless and only gravitationally interacting. A confirmed non-gravitational interaction with neutrinos would mean dark matter has richer microphysics (new forces or mediators) Ś with huge implications for particle-physics model building and laboratory searches.
It links particle physics and cosmic structure. Cosmological surveys would become direct probes of particle interactions in the dark sector, turning maps of galaxies and weak lensing into particle detectors on cosmic scales.
It resolves a concrete cosmological headache. The S₈ tension is one of several small but persistent discrepancies between early-universe (CMB) and late-universe (lensing, galaxy clustering) measurements. A physically motivated resolution reduces the need to invoke systematic errors or contrived fixes.
How the physics produces the effect
Neutrinos stream through the early Universe; dark matter seeds structure. If neutrinos scatter off dark matter weakly, they act like a drag term that suppresses the growth of small-scale structure at late times. That suppression shows up in the matter-power spectrum and in weak-lensing measurements, creating exactly the kind of reduction in clustering amplitude that the S₈ tension describes. The paper quantifies the required interaction strength and shows it can match recent ACT / DES Y3 / SDSS lensing signals when included in the cosmological model.
Papers describing high-throughput cryo-ET pipelines that combine FIB-milling sample preparation, automated tilt-series collection, and large-scale processing Ś prerequisites for building population-level cellular atlases.
Important caveats Ś why we shouldnÆt pop champagne yet
Statistical evidence is suggestive, not definitive. Current analyses find a hint at the ~2¢3σ level depending on datasets and priors Ś interesting and worthy of follow-up, but below the threshold weÆd call a discovery. More data and independent analyses are essential.
Degeneracies and systematics matter. Astrophysical processes (baryonic feedback, galaxy¢halo modeling) and choices in how lensing systematics are handled can mimic or mask this effect; disentangling those possibilities is nontrivial.
Model dependence. ōDM¢neutrino interactionö is a phenomenological description; different microphysical realizations (massive mediators, resonant scattering windows, velocity-dependent cross sections) have different signatures. The next step is to map cosmological hints to concrete particle models and lab tests.
How this will be tested next (fast)
Near-term data rechecks. Teams will reanalyse ACT, Planck, DES Y3 and KiDS data with alternative systematics treatments to test robustness.
Upcoming surveys will be decisive. Rubin Observatory/LSST, ESAÆs Euclid, and next-generation CMB experiments (CMB-S4, Simons Observatory) will map structure and lensing with much higher precision and can confirm or rule out the tiny suppression pattern predicted by DM¢neutrino coupling. Forecast studies show weak-lensing from Rubin/LSST will be especially powerful for these tests.
Cross-discipline constraints. Particle-physics experiments, neutrino observatories, and direct/indirect dark-matter searches will try to bound or detect the mediators implied by the cosmological fits Ś giving a path from sky maps to the lab.
Conclusion
This is the kind of incremental but concrete result that turns cosmology from a descriptive account of the Universe into a precision tool for new particle physics. If real, the DM¢neutrino interaction hypothesis would (1) provide a testable extension of ΛCDM, (2) stimulate new laboratory searches for dark-sector mediators, and (3) reorient strategy for future surveys and simulation campaigns. For research consultants, funders, and biotech/tech sectors that pay attention to deep-tech spinouts, itÆs a reminder that ōfundamentalö discoveries can have long downstream effects in instrumentation, sensors, and data science.
References
1. Zu, L. et al.A solution to the S₈ tension through neutrino¢dark matter interactions.Arxiv. 2026.A solution to the S8 tension through neutrino¢dark matter interactions
2. JZu, L., GiarĶ, W., Zhang, C. et al. Nat Astron (2026). A solution to the S8 tension through neutrino¢dark matter interactions. DOI: 10.1038/s41550-025-02733-1 A solution to the S8 tension through neutrino¢dark matter interactions
3. University of Sheffield press release summarizing the results and data used (ACT, DES, Planck).University of Sheffield. 2026 Scientists find evidence dark matter and neutrinos may interact, challenging standard model of the universe.Scientists find evidence dark matter and neutrinos may interact, challenging standard model of the universe
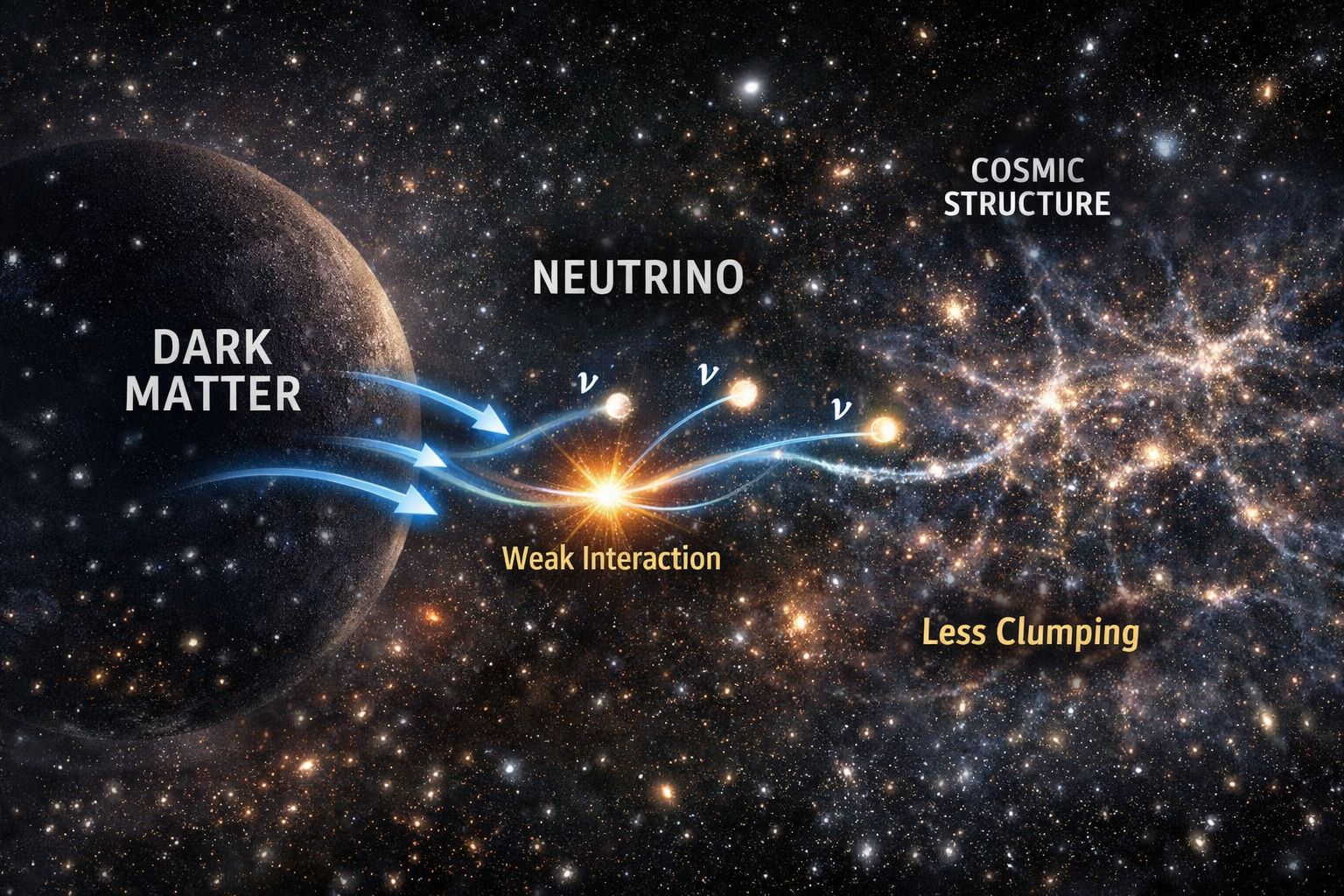
Back to top
Quantum Echoes: How GoogleÆs New Algorithm Brings Quantum Computing Closer to Real-World Applications
By Pedro Paulo Gattai Gomes, Ph.D.
In recent months of 2025 we witnessed a milestone that many in the scientific community considered the next major step: a quantum algorithm that not only outperforms classical computers in speed for a concrete task, but whose output is verifiable Ś meaning other systems can confirm the result is correct. The work, led by Google Quantum AI and named ōQuantum Echoes,ö was described in detail by the team and covered in both scientific and mainstream press as an advance toward a ōverifiable quantum advantage.ö
What ōQuantum Echoesö is Ś in plain terms
Quantum Echoes exploits a protocol of reversible time evolution: the quantum system is evolved forward (a ōgoö), a controlled perturbation is applied, and then the system is evolved backward (a ōreturnö) to recover an amplified signal Ś the ōechoö Ś that encodes information about complex correlations in the system. This trick enables extraction of many-body quantum properties that would be extremely costly to simulate on a classical supercomputer. In the announced experiments, the protocol ran on the Willow processor (≈100+ qubits) and produced results that, according to the authors, would be impractical to reproduce with classical methods within a reasonable timeframe.
Why this matters (and why itÆs more than ōa recordö)
There are three practical reasons the work matters beyond a technical benchmark:
1. Verifiability. Many earlier demonstrations of ōquantum advantageö showed speedups but left doubts about result reliability or practical usefulness; here, the authors emphasize that the result can be verified by alternative experiments (or on different quantum hardware), reducing the risk of experimental artifacts.
2. Applications in chemistry and spectroscopy. The team demonstrated an experimental use case aimed at nuclear magnetic resonance (NMR) applied to model molecules, showing the technique can recover structural information relevant for drug discovery and materials science Ś areas where faithful representations of quantum dynamics are crucial. This is a concrete step toward scientific and industrial applications.
3. A route to verifiable quantum datasets. The approach also opens the possibility of generating datasets produced by quantum systems (with cross-verification) to train AI models or characterize materials that challenge classical simulation Ś a new type of scientific input.
What the numbers say (without sensationalism)
Journalistic coverage quoted striking orders of magnitude Ś for example, comparisons stating that the quantum hardware solved tasks that would take years on classical supercomputers Ś but itÆs important to interpret those numbers carefully: the advantage depends on the specific problem, the comparison assumptions, and the state of the art of the classical baseline algorithms. Put simply: this is a convincing advance for a family of important problems, but it does not mean everyday problems will suddenly be solved instantly by quantum computers.
Limitations and next steps
Even with warranted enthusiasm, clear limitations remain:
Conclusion
For readers interested in applied science, the Quantum Echoes announcement deserves highlighting as a pragmatic milestone: not an ōendö for classical computing nor an immediate promise of large-scale commercial applications, but a technical advance that narrows the gap between lab demonstrations and real scientific problems (specifically in quantum system modeling and spectroscopic analysis). Over the next 12¢36 months, expect attempts at reproduction, adaptations of the protocol to more complex chemical problems, and potential partnerships between research centers and the pharmaceutical/materials industry to explore early use cases.
References
1. Gibney E.Google claims æquantum advantageÆ again Ś but researchers are scepticalNature. 2025.Google claims æquantum advantageÆ again Ś but researchers are sceptical
2. Mi, X., Kechedzi, K. Google Research (2025). A verifiable quantum advantage. A verifiable quantum advantage.
3. Zhang, C., et al.PDF of the technical article/report with experimental details and NMR applications. 2025.Quantum computation of molecular geometry via many-body nuclear spin echoes

Back to top
A patient-specific base-editing therapy for a neonate: a watershed moment for in-vivo gene editing
By Pedro Paulo Gattai Gomes, Ph.D.
In 2025 a multidisciplinary team delivered a personalized, in-vivo CRISPR base-editing therapy to an infant with severe carbamoyl-phosphate synthetase 1 (CPS1) deficiency, producing rapid clinical improvement and demonstrating that individualized, systemically delivered base editing can be safe and therapeutically effective in a human patient. This case is not the end of the story, but it marks the first credible demonstration that tailored, single-patient gene-editing interventions can be executed clinically and supervised with established safety follow-up frameworks.
Introduction - Why its matter?
For nearly two decades the gene-editing field has matured from conceptual tools to engineered nucleases and then to refined base and prime editors that can change single nucleotides without inducing double-strand breaks. Yet translating those tools into safe, systemic human treatments has remained a major bottleneck because of delivery barriers, off-target risk, immunogenicity, and regulatory constraints. The 2025 report of a patient-specific, in-vivo base-editing intervention (delivered via lipid nanoparticles to the liver) that corrected a pathogenic variant in CPS1 is therefore a pivotal demonstration: it shows that the full pipeline Ś rapid design of an individualized editor, GMP-compatible manufacturing of a delivery formulation, and clinically-supervised systemic dosing Ś can be done in a time frame and with a safety profile acceptable for an infant with a life-threatening disorder.
The case and the intervention
A neonate diagnosed with severe CPS1 deficiency received an intravenous, patient-tailored base-editing therapy targeted to the causal variant in hepatocytes. The therapeutic strategy used a base editor (which chemically converts one base to another without creating a double-strand DNA break) packaged into lipid nanoparticles to achieve liver tropism. Early clinical readouts showed marked biochemical and clinical improvement, and no acute safety signals were reported during the initial observation window. The work was published in a peer-reviewed case report in the New England Journal of Medicine and accompanied by institutional press releases from the treating centers.
Technical assessment
1. Choice of editing modality. Base editors are well suited to correcting many point mutations because they make single-base changes without requiring double-strand breaks, reducing the probability of large indels or translocations. The selection of a base editor for a neonatal metabolic disorder, where hepatocytes are the therapeutic target, is mechanistically appropriate Ś provided that guide design, editing window, and cellular context align to minimize bystander edits. The authors presented design choices and in-vitro validation data supporting the specificity of the chosen editor.
2. Delivery strategy. Systemic lipid nanoparticle (LNP) delivery to the liver is now a proven route (recall successful siRNA and mRNA therapeutics). LNPs offer transient exposure of the editor machinery, which reduces persistent expression windows and theoretically narrows the period during which off-target editing can occur. The team leveraged this established delivery modality and prior translational experience to reach therapeutic hepatocyte editing levels.
3. Timeframe from genotype to therapy. Perhaps the most striking operational achievement is the rapid turnaround Ś designing a bespoke therapeutic within months Ś which required tight collaboration among clinicians, molecular engineers, and GMP manufacturing. This rapid customization is central to the promise of personalized therapies for ultra-rare pediatric diseases, but it also raises questions about scalability and regulatory pathways if many such individualized therapies are to be produced routinely.
4. Safety and monitoring.The initial safety dataset is reassuring but necessarily limited: a small number of days to months of follow-up can detect acute toxicity, infusion reactions, and some early molecular signals, yet cannot exclude late-onset outcomes (e.g., oncogenic events, delayed immune responses). The treating teams have committed to long-term monitoring, consistent with emerging guidance for genome-editing therapeutics.
Broader implications for research and clinical translation
Limitations and sober caveats
Where we go from here
1. Standardized preclinical pipelines for rapid safety and off-target profiling that can be used across custom designs.
2. Validated manufacturing platforms that reduce per-patient costs and cycle times while ensuring reproducibility.
3. Longitudinal registries and shared datasets so single-patient cases can be contextualized and aggregated to detect rare adverse events.
4. Ethics frameworks and equitable access models so personalized editing does not become a privilege limited to the few.
Conclusion
As a researcher who has followed genome engineering from early ZFNs through CRISPR and now to base and prime editors, I view this 2025 personalized base-editing intervention not as a shimmering finish line but as a responsibly optimistic milestone. It demonstrates the technical feasibility of a pipeline that had long been a theoretical possibility: take a defined pathogenic variant, design a precise editor, deliver it systemically, and observe clinical benefit. That pipeline will not be trivial to scale, nor devoid of risk, but the successful case provides a proof of principle that will Ś and should Ś accelerate careful, data-driven translational work for other life-threatening monogenic diseases.
References
1. Musunuru K., et al.Patient-Specific In Vivo Gene Editing to Treat a Rare Genetic Disease.N Engl J Med. 2025. DOI: 10.1056/NEJMoa2504747. Patient-Specific In Vivo Gene Editing to Treat a Rare Genetic Disease
2. ChildrenÆs Hospital of Philadelphia / Penn Medicine. WorldÆs First Patient Treated with Personalized CRISPR Gene-Editing Therapy (press release). May 2025. World's First Patient Treated with Personalized CRISPR Gene Editing Therapy at ChildrenÆs Hospital of Philadelphia
3. National Institutes of Health (NIH) News Release.Infant with rare, incurable disease is first to successfully receive personalized gene-therapy treatment. May 15, 2025.Infant with rare, incurable disease is first to successfully receive personalized gene therapy treatment
4. Nature / Science commentary and press coverage (summaries and independent expert perspectives) discussing the broader implications of the case and placing it in context of gene-editing clinical progress in 2025.CRISPR used for the first time to treat a rare metabolic disease in a baby
5. Reviews of CRISPR clinical translation and in-vivo editing technologies (2025) that outline delivery, safety, and regulatory challenges relevant to personalized interventions.Advancing gene editing therapeutics: Clinical trials and innovative delivery systems across diverse diseases
Back to top
EchoBack-CAR: when sound becomes a switch
By Pedro Paulo Gattai Gomes, Ph.D.
In the lab, the best innovations often arise at the intersection of a well-known practical limitation and a clever engineering idea that overcomes it. ThatÆs the case with whatÆs been called EchoBack-CAR Ś a platform for sonogenetic CAR-T therapy that uses focused ultrasound (FUS) to remotely and locally activate expression of a chimeric antigen receptor, showing marked gains against solid tumors and reduced off-target effects. Reading the primary studies and critical reviews shows this is a plausible and useful technological advance, but one that requires caution when extrapolating to clinical use.
The technical gist Ś how ōsoundö becomes therapy
The idea is elegant in its operational simplicity. T cells are genetically engineered with a circuit placing the CAR gene under control of a heat-sensitive promoter (heat-shock element, HSE). Applying focused ultrasound to the tumor volume produces a localized mild temperature increase (typically 39¢43 ░C) that activates the promoter, drives robust CAR expression, and triggers cytotoxic activity only where itÆs needed. To prolong and amplify the effect after a brief pulse, the authors integrated a positive feedback loop: initial CAR signaling reinforces the transgeneÆs own expression, producing windows of activity much longer than those achieved with conventional FUS-inducible systems. This strategy Ś demonstrated in 3D models and mouse studies Ś produced better tumor control than constitutive CARs and showed lower systemic toxicity.
Why this matters now?
Three practical points justify the technical enthusiasm:
1. Realistic spatial precision. FUS can reach deep tissue with millimeter-scale resolution and is already used clinically for ablation and neuromodulation, which simplifies potential translation without inventing new hardware.
2. Mitigation of on-target/off-tumor toxicity. Activating CAR expression only at the tumor site reduces the risk of severe adverse events caused by antigen recognition in normal tissues. Preclinical data show a promising safety profile relative to constitutive CARs.
3. Counteracting antigenic heterogeneity. Programmable FUS pulses enable ōprimingö and sequential reactivation strategies useful for tumors with heterogeneous antigen expression or for controlling relapse. Related work also explores FUS to induce expression of candidate antigens in tumors, increasing their immune visibility.
Primary evidence and supporting articles
The core experimental work formalizing EchoBack is described in a detailed paper (a series of in-vitro experiments, 3D organoids, and murine models) and is accompanied by translational reports and methodological reviews validating HSEs and FUS as biological triggers. Key references for assessing technical robustness include the EchoBack engineering article, follow-up translational reports, and reviews on FUS-induced transgene control.
Note: mainstream scientific coverage (for example, reporting in major journals) has been measured and tied to these preclinical results; media attention helps map the advance but does not replace independent validation.
Real limitations Ś whatÆs still needed before clinical use
As a researcher experienced in biomedical translation, I stress essential points that any clinical development plan must address:
Practical outlook (12¢36 months)
If preclinical evidence holds and industry interest emerges, I expect:
1. GLP toxicology studies and proof-of-concept trials in larger animals using clinical FUS devices.
2. Studies mapping the ōactivation windowö across thermal doses and pulse protocols to optimize dose/time parameters.
3.Industry-academic partnerships to produce EchoBack cells under GMP and to validate combined FUS-cell therapy workflows.
Conclusion Ś practical prudence, technical optimism
EchoBack-CAR exemplifies the direction immunotherapy should take: bold on longstanding problems (antigen scarcity, on-target toxicity, exhaustion) and disciplined on engineering (sensitive promoters, synthetic feedback, controlled delivery). ItÆs a conceptual and technological advance that deserves attention from scientists, regulators, and clinical engineers Ś but it is not yet a clinical-mature solution. As always, the path from bench to bedside demands rigorous replication, independent validation, and thorough safety testing.
References
1. Longwei, Liu, et al.Engineering sonogenetic EchoBack-CAR T cells.Cell. 2025. DOI: 10.1016/j.cell.2025.02.035. Epub 2025 Apr 2. Engineering sonogenetic EchoBack-CAR T cells
2. Jeungyoon, Lee et al.An integrated ultrasound-guided focused ultrasound system enables spatiotemporal control of thermal gene activation in engineered immune cells 2025. DOI: 10.7150/thno.118405. An integrated ultrasound-guided focused ultrasound system enables spatiotemporal control of thermal gene activation in engineered immune cells
3. Yoon, CWTumour priming by ultrasound mechanogenetics for CAR T therapy. Feb, 2025.DOI: 10.1038/s41563-025-02391-8. Epub 2025 Oct 31.Tumour priming by ultrasound mechanogenetics for CAR T therapy
4. Yiqian, Wu.Ultrasound Control of Genomic Regulatory Toolboxes for Cancer Immunotherapy. 2025.Ultrasound Control of Genomic Regulatory Toolboxes for Cancer Immunotherapy.
5. Gunjan, Dagar.Reviews of CRISPR clinical translation and in-vivo editing technologies (2025) DOI: 10.1186/s12967-023-04292-3Harnessing the potential of CAR-T cell therapy: progress, challenges, and future directions in hematological and solid tumor treatments
Back to top
A Genetic Switch for Heat-Resilient Rice: Why a 2025 Cell Study Matters for Food Security
By Pedro Paulo Gattai Gomes, Ph.D.
In the middle of a rapidly warming century, one of the most consequential scientific questions is not whether crops will face more heat stress, but how far biology can be pushed to adapt. Rice sits at the center of that problem because it feeds billions, yet its reproductive stage is unusually vulnerable to warm nights and heat waves. A classic PNAS study showed that rice yield declined by about 10% for every 1░C increase in growing-season minimum temperature in the dry season, providing an early and very clear signal that nighttime warming is not a minor stressor but a direct agronomic threat.
A strikingly recent advance came in 2025, when a Cell paper reported a natural genetic system that improves both grain quality and yield under field heat stress. The authors evaluated a rice mini-core collection of 533 accessions across seasons and locations, then identified a major locus, QT12, that acted as a negative regulator of grain-quality thermotolerance. What makes this study especially compelling is that it was done in real field conditions, not only in controlled chambers, which gives the result unusual practical weight.
At the mechanistic level, the study is even more interesting than the headline. QT12 was shown to disrupt endosperm storage-substance homeostasis by over-activating the unfolded protein response, linking heat exposure to the deterioration of grain quality. The paper also identified a natural regulatory circuit involving NF-Y transcription factors: under high temperature, interactions among NF-YB9, NF-YC10, and NF-YA8 weaken, which releases repression of QT12 and allows the heat-sensitive phenotype to emerge. In other words, the plant is not simply ōdamaged by heatö; it is decoding heat through a specific gene-regulatory architecture.
The practical significance is hard to overstate. In large-scale high-temperature trials, low QT12 expression was associated with superior grain quality and an increase in elite rice yield of up to 1.31¢1.93-fold. The authors also reported natural variations that function as an on-off switch, opening a realistic path for conventional breeding as well as gene editing. For a crop where breeders have often had to trade yield against quality, this is exactly the kind of result that can reshape strategy rather than merely refine it.
This 2025 discovery did not appear in isolation. Another 2025 Cell Reports study identified the OsEBF1-OsEIL5-OsPP91 module as a regulator of rice heat tolerance through ubiquitination and transcriptional activation, reinforcing the idea that thermotolerance is controlled by layered signaling networks rather than a single master gene. A separate 2025 Cell paper went further upstream, showing that plant heat sensing can begin with membrane lipid remodeling and proceed through a nuclear signaling cascade. Taken together, these studies suggest that heat resilience in plants is being uncovered as a multilevel system: membrane, protein stability, transcription, and field phenotype are all connected.
That broader perspective matters because scientific breakthroughs are most useful when they dissolve old false choices. For decades, crop improvement under heat was often framed as a compromise between productivity and quality, or between laboratory elegance and field relevance. The QT12 work challenges both assumptions. It shows that a naturally occurring allele can preserve grain quality while sustaining yield under real heat stress, and it does so through a mechanism that breeders can actually work with. This is the difference between an interesting molecular finding and a discovery with translational force.
Of course, no responsible scientist should oversell a single paper. Field validation across more environments, genetic backgrounds, and seasons will still be essential, and any deployment strategy must be tested against the complexities of local agronomy. But that caution does not diminish the importance of the discovery; it strengthens it. The best science does not merely announce a result. It changes the questions that serious researchers are now obliged to ask. In this case, the question is no longer whether rice can be genetically tuned for heat tolerance in the field. It is how quickly that knowledge can be translated into varieties that remain productive in a hotter world.
References
1. Li, W. et al.A natural gene on-off system confers field thermotolerance for grain quality and yield in rice.Cell (2025). DOI:10.1016/j.cell.2025.04.011. A natural gene on-off system confers field thermotolerance for grain quality and yield in rice
2. Peng, S. et al.Rice yields decline with higher night temperature from global warming. PNAS (2004). DOI:10.1073/pnas.0403720101 Rice yields decline with higher night temperature from global warming
3. Liu, J. et al.The OsEBF1-OsEIL5-OsPP91 module regulates rice heat tolerance via ubiquitination and transcriptional activation. Cell Reports (2025). DOI: 10.1016/j.celrep.2025.115271. The OsEBF1-OsEIL5-OsPP91 module regulates rice heat tolerance via ubiquitination and transcriptional activation
4. Kan, Y. et al..A stepwise decoding mechanism for heat sensing in plants connects lipid remodeling to a nuclear signaling cascade. Cell (2025). A stepwise decoding mechanism for heat sensing in plants connects lipid remodeling to a nuclear signaling cascade

Back to top
TimeVault: turning a mysterious cellular structure into a molecular memory device
By Pedro Paulo Gattai Gomes, Ph.D.
For decades, vault particles have been one of cell biologyÆs most enigmatic structures: huge ribonucleoprotein cages found across most eukaryotic lineages, structurally conserved yet still only partly understood. That mystery is exactly what makes the latest advance so exciting. In a 2026 Science paper, Yu-Kai Chao, Michelle Wu, Qiyu Gong, and Fei Chen describe TimeVault, a genetically encoded system that repurposes engineered vault particles to store transcriptomes inside living mammalian cells for later readout.
At its core, the breakthrough solves a long-standing problem in molecular biology: most transcriptomic measurements are snapshots, even though biology unfolds as a movie. The TimeVault system captures mRNA through poly(A)-binding protein, stores that RNA in vault particles, and preserves the transcriptome in living cells for more than seven days. In the authorsÆ experiments, it enabled high-fidelity recording with minimal cellular perturbation, including the capture of transient stress responses and gene-expression states associated with drug-naive persister cells in lung cancer under EGFR inhibition.
Why does this matter?
Because many important cellular decisions are temporal. Development, stress adaptation, and therapy resistance often depend on brief molecular states that disappear before a standard RNA-seq assay can catch them. TimeVault fits into the broader field of cellular recording, which has long aimed to preserve fleeting biological information in a durable molecular format. What is unusual here is the medium: instead of writing history into DNA, the system stores it in a repurposed intracellular particle.
This is not the first hint that vault biology may be functionally important. Earlier work showed that the vault RNA vtRNA1-1 can influence apoptosis independently of the vault complex itself, suggesting that vault-associated components are not just cellular decoration but active players in gene regulation and survival pathways. More recently, structural studies have helped clarify how the human vault cage is assembled and how PARP4 and NAD+ interact with it, providing a stronger mechanistic foundation for engineering the particle into a useful tool.
The deeper significance of TimeVault is conceptual. It adds a new way to reconstruct cellular history without relying on constant live imaging or destructive sampling. That could make it especially valuable for developmental biology, stress-response mapping, and cancer research, where the most important state may be the one that existed briefly and vanished. In that sense, the discovery is not just about vaults; it is about giving cells a readable memory.
References
1. Chao, Y.-K., Wu, M., Gong, Q., & Chen, F.A genetically encoded device for transcriptome storage in mammalian cells.Science. 2026. DOI: 10.1126/science.adz9353. A genetically encoded device for transcriptome storage in mammalian cells
2. Lodwick, J. E., et al. Structural insights into the roles of PARP4 and NAD+ binding in the human vault cage. Nature Communications. 2025. DOI: 10.1038/s41467-025-61981-x Structural insights into the roles of PARP4 and NAD+ binding in the human vault cage
3. Amort, M., et al.Expression of the vault RNA protects cells from undergoing apoptosis. Nature Communications. 2015. DOI: 10.1038/ncomms8030 Expression of the vault RNA protects cells from undergoing apoptosis
4. Sheth, U., et al. DNA-based memory devices for recording cellular events.PMC Review. 2018. DOI: 10.1038/s41576-018-0052-8 DNA-based memory devices for recording cellular events
Back to top
The Hidden Architecture of Centromeres: Why a 2025 Epigenetic Discovery Matters
By Pedro Paulo Gattai Gomes, Ph.D.
For many years, centromeres were treated as one of the genomeÆs most stubborn paradoxes: essential for chromosome inheritance, yet still resistant to a fully satisfying molecular explanation. We knew that CENP-A marks centromeric chromatin, and we knew that DNA sequence alone could not account for centromere identity. But what has become increasingly clear in the last few years is that centromeres are not simply ōplacedö by sequence or one epigenetic mark. They are actively maintained by a layered chromatin environment, and a pair of 2025 studies has sharpened that view in an especially important way.
The first study, published in Nature Genetics on 4 September 2025, showed that DNA methylation is not a passive passenger at human centromeres: it causally influences CENP-A positioning and centromere function. When the authors induced rapid centromeric demethylation, they observed altered centromeric protein binding, changes in centromere architecture, aneuploidy, and reduced cell viability. In other words, centromeric DNA methylation is not merely correlated with centromere integrity; it helps determine where the centromere lives and how well it works.
That finding already would have been enough to shift the conversation. But a second paper, published in Nature Structural & Molecular Biology on 25 November 2025, took the story one step deeper. Using single-molecule epigenomics, the authors found that heterochromatin boundaries help maintain centromere position, size, and number. They implicated H3K9me3 methyltransferases SUV39H1/H2 and SETDB1, as well as noncanonical roles for SUZ12, in preserving those boundaries. When those boundaries fail, the primary CENP-A domain can expand, shift, or even seed additional functional CENP-A domains on the same alpha-satellite sequence.
Taken together, these papers do something more important than add detail to an old model. They move centromere biology away from a simple ōmarker-centricö view and toward a territorial view of chromatin. A centromere is not just a spot that contains CENP-A. It is a controlled chromatin landscape, bounded by methylation, shaped by heterochromatin, and stabilized by enzymes that preserve domain architecture. That shift matters because chromosome segregation is only as faithful as the epigenetic boundaries that support it.
The broader implication is hard to overstate. If centromere identity can be destabilized by changes in DNA methylation and heterochromatin boundary control, then centromere dysfunction is not merely a structural curiosity. It becomes a plausible route into aneuploidy, genome instability, and disease biology. The Nature Genetics paper explicitly links centromeric hypomethylation to chromosome mis-segregation and loss of viability, while the NSMB study shows how boundary erosion permits ectopic centromere nucleation. That is a mechanistic bridge from chromatin chemistry to cellular fate.
There is also a conceptual elegance here that seasoned molecular biologists tend to appreciate. Centromeres have long been described as epigenetic, but epigenetic is not a synonym for vague or unstable. These studies show precision: a particular methylation landscape, a particular heterochromatin boundary system, and a particular chromatin domain size together preserve centromere identity. That is the kind of result that advances a field, because it converts a general principle into a testable molecular architecture.
For researchers, the next questions are now much sharper. Which lesions in centromeric methylation are tolerated, and which cross the threshold into instability? Which diseases reflect cause rather than consequence? And can centromere boundary maintenance be manipulated therapeutically, especially in contexts where chromosomal instability is a defining feature? The recent papers do not answer those questions, but they make the questions far more precise than before.
References
1. Salinas-Luypaert, C. et al.DNA methylation influences human centromere positioning and function.Nature Genetics (2025). DOI: 10.1038/s41588-025-02324-w. DNA methylation influences human centromere positioning and function
2. Carty, B. L.et al. Heterochromatin boundaries maintain centromere position, size and number Nat Struct Mol Biol. 2026. DOI: 10.1038/s41594-025-01706-2 Heterochromatin boundaries maintain centromere position, size and number
3. Naughton, C. et al.Human centromere repositioning activates transcription and opens chromatin fibre structure. Nature Communications (2022) DOI: 10.1038/s41467-022-33426-2 Human centromere repositioning activates transcription and opens chromatin fibre structure

Back to top
Challenging a Central Dogma of DNA Repair
By Pedro Paulo Gattai Gomes, Ph.D.
For decades, we have operated under the well-established principle that cells rely on two primary, DNA-dependent mechanisms to repair the most dangerous type of genomic insult: the DNA double-strand break (DSB). These canonical pathways are homologous recombination (HR), an elegant but slow high-fidelity process restricted to the S/G2 phases of the cell cycle, and non-homologous end joining (NHEJ), a faster but more error-prone process that can act throughout the cell cycle. The underlying assumption was always that the template for repair, whether it be a sister chromatid or the broken end itself, must be DNA.
This new study elegantly shatters that assumption. The researchers provide definitive evidence for a novel, alternative pathway termed RNA-templated double-strand break repair (RT-DSBR). In this process, a cellular messenger RNA (mRNA) transcriptŚa copy of genetic information that we traditionally thought of solely as a protein-coding intermediaryŚcan be physically used as a scaffold and template to guide the precise repair of a broken DNA molecule. The work challenges long-held views about the functional interplay between RNA and DNA, showing that the flow of information in the cell can, in times of crisis, be reversed and repurposed to preserve genomic integrity.
The Molecular Cast: Polζ as an Unconventional Reverse Transcriptase
A key element of this discovery lies in the identification of the molecular machinery responsible for this feat. To copy the RNA template back into DNA, the cell requires an enzyme with reverse transcriptase (RT) activity. Unlike retroviruses that encode their own dedicated RT, human cells do not. Through a meticulous CRISPR/Cas9-based genetic screen, the team identified a surprising candidate: DNA polymerase ζ (Polζ).
Polζ is a specialized translesion synthesis polymerase previously known for its role in bypassing DNA lesions that block the main replicative polymerases. That this same enzyme possesses robust reverse transcriptase activityŚthe ability to synthesize a DNA strand using an RNA templateŚwas an unexpected and significant finding. It reveals a remarkable functional plasticity in our cellular enzymatic toolkit, where a polymerase we thought we understood has been moonlighting in a completely different, and critical, role.
Evidence and Implications: From the Lab Bench to the Cancer Genome
The evidence presented is both elegant and robust. The authors developed a clever suite of fluorescence-based and sequencing-based reporter assays to definitively track and quantify RT-DSBR events in living human cells. These assays showed that both synthetic RNA oligonucleotides and naturally occurring mRNAs could direct the repair of a Cas9-induced break.
Crucially, the study connects this molecular mechanism directly to human disease. By analyzing large-scale cancer genome sequencing datasets, the researchers uncovered a distinct mutational signature: whole intron deletions (WIDs). These are precise deletions of entire intronic sequences from a gene, a tell-tale sign that a spliced mRNA (which naturally lacks introns) served as the repair template. The presence of WIDs in various cancer genomes provides compelling evidence that RT-DSBR is activeĀin vivoĀand contributes to the landscape of somatic mutations that drive tumor evolution.
A New Frontier in Genomic Research and Therapeutic Targeting
The implications of this work extend far beyond a single publication. The discovery of the RT-DSBR pathway opens up a host of new research avenues. It forces us to reconsider the full repertoire of DNA repair options available to a cell, particularly in transcriptionally active regions where RNA templates are abundant. The fact that this pathway can lead to characteristic mutations like WIDs also provides a new lens through which to interpret the complex mutational signatures seen in cancers.
From a therapeutic perspective, Polζ and its associated protein partners may now become targets of interest in oncology. In tumors that are deficient in HR (such as those with BRCA1/2 mutations), these cells become exquisitely dependent on alternative repair pathways for survival. Targeting RT-DSBR by inhibiting Polζ could represent a novel synthetic lethal strategy to selectively kill such cancers. Furthermore, the discovery presents both opportunities and cautionary notes for the gene-editing field. While RNA templates could be engineered for precise genome modifications, the potential for off-target or unintended RT-DSBR events during CRISPR-based editing must now be carefully considered.
A Signpost for Future Inquiry
In over 17 years of watching molecular biology unfold, you learn to appreciate the moments when a field takes a real step forward. The characterization of RNA-templated DSB repair is one of those moments. It is a reminder that our fundamental understanding of cellular machinery is never truly complete, and that the most elegant solutions are often hiding in plain sight. For researchers, it is a clear signpost pointing toward the next generation of questions about genome stability, evolution, and disease. For the broader community, it is a powerful illustration of the enduring importance of basic, curiosity-driven science to illuminate the hidden workings of life and pave the way for future medical advances.
References
1. Jalan et al.ĀRNA transcripts serve as a template for double-strand break repair in human cells.Nature Communications (2025). DOI: 10.1038/s41467-025-59510-x. RNA transcripts serve as a template for double-strand break repair in human cells
2. American Association for Cancer Research RNA to the Rescue: A New DNA Repair Pathway for Double-Strand Breaks Accessed on 2026. RNA to the Rescue: A New DNA Repair Pathway for Double-Strand Breaks
>

Back to top

Leave a Comment
Comments: